Division of Translational Pediatric Sarcoma Research
Prof. Dr. Dr. Thomas Grünewald
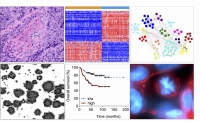
Translational spectrum of methods: histology, omics-analyses, preclinical models, correlation with clinical data
© dkfz.de
The mission of the division of Translational Pediatric Sarcomas Research, which corresponds to the KiTZ division of the same name, is to improve treatment options for children and adolescents affected by sarcomas. We aim at uncovering disease mechanisms that can be used diagnostically and therapeutically to improve the long-term chances of recovery of our young patients. The focus lies on new methods that are essential for correct diagnosis and the choice of the best possible therapy. We also investigate less aggressive therapies and new approaches to overcome drug resistance of tumors. In this regard, one special objective is to decipher the interactions between acquired mutations and innate natural variants of the genome, which can play a decisive role in the development and progression of cancer, particularly in Ewing sarcoma.
We aim at developing novel therapeutic strategies based on new insights into the molecular mechanisms of disease following two major lines of research (see research areas). Our investigations start with the analysis of the patients tumors in situ, followed by studying patient-derived cell lines using a wide variety of in silico, in vitro, and in vivo techniques.
The division is supported by the Barbara und Wilfried Mohr Stiftung.
Research areas
Tumorigenesis in the context of molecular signaling pathways of child development
Certain oncogenes often hijack or interfere with molecular signaling pathways important for child development, thereby conferring a considerable growth advantage to tumor cells. We explore the mechanisms through which oncogenes arrest sarcoma cells in a specific but yet largely undifferentiated state via deregulation of key developmental pathways, and how they cooperate with them to promote tumorigenesis and cancer progression.
Therapy resistance, tumor heterogeneity, and predictive biomarkers
Primary and/or secondary resistance to conventional chemotherapeutic drugs is a frequent event in pediatric sarcoma. Chemotherapy-resistance is a highly specific process in which tumors become resistant to certain drugs while maintaining (limited) susceptibility toward others. Underlying to this resistance is considerable plasticity due to intra-tumor heterogeneity, which enables the adaptation to therapeutic cell stress on the clonal and sub-clonal level. Therefore, we strive for illuminating the (epi)genetic basics and biological mechanisms of tumor-heterogeneity in order to identify new biomarkers for individualized risk-stratification. By means of these biomarkers we can predict the response to certain treatments and better assess which targeted therapeutics may help to overcome resistance to conventional (chemo)therapeutics.
The above-mentioned research areas are addressed by three highly synergistic teams within my division of Translational Pediatric Sarcoma Research:
The team Translational Genomics (Prof. T. Grünewald) systematically establishes multi-dimensional omics-datasets of a large number of pediatric sarcomas and correlates molecular alterations with clinical data to identify new driver mutations, disease-relevant drug targets and prognostic/predictive biomarkers, and to create resources for hypothesis generation and functional validation for future projects.
The team Mechanisms of Cancer Progression (Dr. F. Cidre-Aranaz) aims at deconvoluting the multilayered process underlying cancer progression and metastasis. For this purpose, systems biology approaches based on multi-omics data and clinical information with functional in vitro and in vivo experiments will be combined to identify treatable vulnerabilities in pediatric sarcomas that can prevent metastasis.
The team Innovative Therapies (Dr. S. Ohmura) is investigating potential therapeutic targets for pediatric sarcomas in preclinical models. The focus here lies in developing and testing new targeted therapy approaches that enable more effective therapy with fewer side effects, especially for patients with chemo resistant tumors.

