Brain Mosaicism and Tumorigenesis
- Functional and Structural Genomics
- Junior Research Group

Dr. Pei-Chi Wei
Principal Investigator
Not all brain cells have the same DNA. Some of these changes happen early in life and could play a role in the development of diseases like cancer. Our lab studies how these changes in DNA arise, how they are passed on to other cells, and how they spread in the brain.

Our Mission: Unraveling the Role of DNA Diversity in Brain Health
Brain Development is a Dynamic Process
The human brain is built from billions of neurons and astrocytes, generated through hundreds of divisions by neural progenitor cells. Remarkably, most of these cells form within just 24 weeks during fetal development. This process is carefully orchestrated in both time and space, shaping the ~400g newborn brain.
My lab explores fundamental questions: How do neural progenitor cells know when to stop dividing? Why and how do they extend their cell cycles? Does rapid cell division increase DNA damage in these cells? Understanding these mechanisms is key to uncovering how brain cancer begins, progresses, and evolves.
Methods and technologies
We have developed experimental and analytical tools to uncover the interactions between the cellular machinery responsible for copying and reading DNA sequences. Using high-throughput sequencing technologies, we measure the timing and speed of DNA replication and examine the transcriptional "traffic" created by the machinery decoding DNA.
Additionally, we create mouse models to investigate how neural stem cells compete for space and resources during early brain development. Our work also includes designing experimental systems that allow us to "travel back in time," tracing early developmental neural stem cell behaviors at later stages of life.
Goals and societal relevance
Brain cancer prevention faces significant challenges due to our limited understanding of how early cellular processes contribute to tumor formation. Our mission is to bridge this gap by uncovering how neural stem and progenitor cells drive brain development while maintaining DNA integrity. By studying the mechanisms of clonal expansion, cellular competition, and genomic stability, we aim to identify key risk factors and develop strategies for preventing brain cancers. Through this work, we strive to advance brain health and reduce the societal impact of neurological diseases.
Research Projects

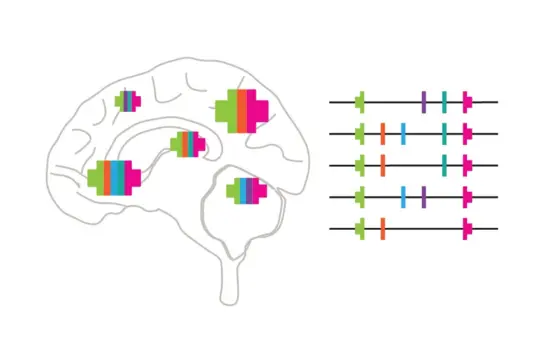
Building a brain takes countless rounds of cell division, and every division has to faithfully copy three billion letters of our genetic code. Sometimes the copying machinery stalls, and the long string of DNA snaps. When the cell glues the broken ends back together in the wrong way, whole chunks of the genetic instructions go missing or get duplicated by accident — unplanned losses and gains the cell never scheduled. These changes are among the strongest known risk factors for autism and brain cancer. Yet why they keep happening at the same spots has remained a puzzle.
We identified a fundamental source. Working with the stem cells that build the brain, we found a handful of "weak spots" in the genetic code. These sit inside the longest genes used by brain cells — the very genes that help neurons connect and talk to each other — and they snap especially easily when DNA copying runs into trouble.
We showed that these weak spots are the starting point for the unplanned losses. The breaking itself is predictable — it happens at the same spots every time DNA copying is stressed — but the repair is not: some cells heal cleanly, some lose the same chunk again and again, forming the recurrent changes seen in disease, and others patch the damage into large, one-of-a-kind rearrangements, leaving each neuronal cell with a slightly different set of genetic building blocks. The same scheduled event is "bad luck" for cells whose repair goes wrong, and a source of natural diversity for the rest — a cell-to-cell variation now recognized as a normal feature of healthy brains, and a contributor to diseased ones. Strikingly, silencing the long genes shut down the breaks and the unplanned changes altogether.
The findings recast these weak spots as a central source of genetic diversity in the developing brain, opening new avenues for understanding how brain disorders and brain tumors begin.
Sources:
Corazzi and Ing et al., (2026) Nature Communications
Corazzi and Ionasz et al., (2024) Nature Communuications
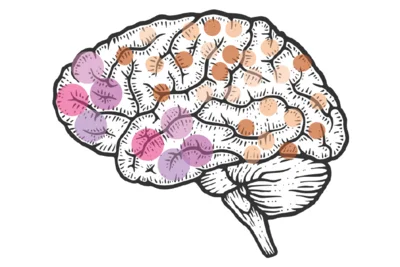
Our research delves into how the size of neural stem cell clones impacts the brain’s structure and function. Neural stem clones are groups of neurons that share the same DNA blueprint, and their size directly influences the spatial organization and functional diversity of neurons in the brain. Intriguingly, studies show that the size of neuronal clones varies across brain regions, suggesting that clone size plays a key role in tailoring the brain’s architecture to its specific needs.
My lab develops preclinical tools and models to investigate clonal expansion, selection, and evolution in the developing brain. We focus on identifying the triggers of clonal competition—a process that can drive unregulated, excessive cell proliferation, also known as precancerous hyperproliferation.
Our goal is to determine whether such unchecked proliferation in normal tissue increases the risk of cancer development. Ultimately, this research could uncover new risk factors and inform strategies for preventing brain cancer.
Current team
-

Dr. Pei-Chi Wei
Principal Investigator
-

Lorenzo Corazzi
Ph.D. student
-

Giulia Di Muzio
Ph.D. student
-

Boyu Ding
Ph.D. student
-

Mila Duerink
Masters student (Home Uni: Leiden, Leiden, NL)
-

Marco Giaisi
Lab Manager
-

Dr. Alex Ing
Senior Scientist
-

Hsin-Jui Lu
Ph.D. student
-
Maya Thomas
Masters student (Home uni: Sorbonne, Paris, FR)
Associated members
Michael Allers
M.D. student with Prof. Dr. Dr. Peter Huber (E055)
E-mail: m.allers(at)dkfz-heidelberg.de
Phone: +49 6221 42 3249
Selected Publications
Lorenzo Corazzi, Alex Ing, Eva Benito, Marco Raffaele Cosenza, Patrick Hasenfeld, Thomas Weber, Anna J. M. Marx, Vivien S. Ionasz, Nathan Trausch, Sarah Benedetto, Giulia Di Muzio, Boyu Ding, Jana Berlanda, Marco Giaisi, Nina Claudino, Thomas Höfer, Jan O. Korbel & Pei-Chi Wei
Giulia Di Muzio, Sarah Benedetto, Li-Chin Wang, Lea Weber, Franciscus van der Hoeven, Brittney Armstrong, Hsin-Jui Lu, Jana Berlanda, Verena Körber, Nina Claudino, Michelle Krogemann, Thomas Höfer, Pei-Chi Wei
Lorenzo Corazzi, Vivien S Ionasz, Sergej Andrejev, Li-Chin Wang, Athanasios Vouzas, Marco Giaisi, Giulia Di Muzio, Boyu Ding, Anna J M Marx, Jonas Henkenjohann, Michael M Allers, David M Gilbert, Pei-Chi Wei
Aseda Tena, Yuxiang Zhang, Nia Kyritsis, Anne Devorak, Jeffrey Zurita, Pei-Chi Wei (co-correspondence), and Frederick W. Alt
Wei PC, Chang AN, Kao J, Du Z, Meyers RM, Alt FW, Schwer B
Open positions
Replication stress is a hallmark of cancer. Oncogene overexpression, loss of tumor suppressors, and other yet-unknown factors naturally promote the formation of DNA breaks through replication stress. It has been speculated that this endogenous replication stress not only drives cancer progression but also fuels its evolution. Paradoxically, however, increasing replication stress—by targeting the DNA replication machinery—has also emerged as an effective strategy to selectively kill cancer cells.
Our laboratory is currently seeking a Ph.D. student to investigate the mechanisms underlying this Achilles’ heel of cancer: replication stress. We have developed both computational and experimental tools to dissect, at single-nucleotide resolution, the molecular processes leading to DNA breaks in the genomes of cancer cells.
Apply the Ph.D. position here:
https://jobs.dkfz.de/en/jobs/168135/phd-student-in-cancer-breaksome
We are recruiting a postdoctoral researcher through DKFZ Marie Skłodowska-Curie MasterClasses. Please apply by May 18, 2026.
Get in touch with us