Chromatin Networks

Prof. Dr. Karsten Rippe
We investigate (epi)genome organization and functions in single cells to reveal how deregulated chromatin features promote cancer development, tumor heterogeneity, and therapy resistance.
Image: Chromatin co-accessibility and topology map of a human genome locus (Seufert et al. 2024),
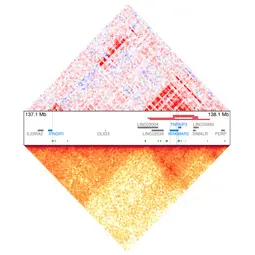

Image: Chromatin co-accessibility and topology map of a human genome locus (Seufert et al. 2024),
Our Research
We integrate single-cell (epi)genomics with advanced fluorescence microscopy methods to understand how chromatin organization and nuclear architecture govern gene regulation and telomere maintenance. Our work combines the imaging-based analysis of cellular phenotypes in their endogenous tissue context with molecular multi-omics profiles that include transcriptome, chromatin accessibility, surface proteins, and immune cell receptors.
This interdisciplinary approach connects genome organization to functional outcomes, revealing how a disrupted nuclear architecture drives aberrant cell states in cancer. We test the mechanisms identified in cell model systems through targeted perturbations, such as light-controlled epigenetic editing. This integrated methodology helps us understand tumor heterogeneity, therapy resistance mechanisms, and the unlimited proliferation potential of cancer cells in both blood cancers and solid tumors.
Bioquant-Website
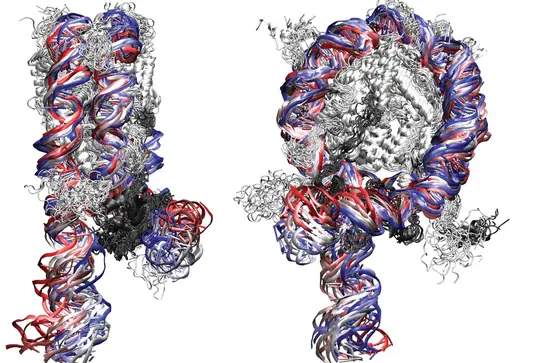
Computer simulations of the dynamic nucleosome structure with the DNA wrapped around the histone protein core (Ettig 2011 Biophys J). The nucleosome is the building block of chromatin. In a human cell about 30 million nucleosomes organize the genome of 6 billion DNA base pairs.
Our Team
-

Prof. Dr. Karsten Rippe
-

Mislav Basic
-

Irene Gerosa
-

Caroline Knotz
-

Emma Koeleman
-

Dr. Norbert Mücke
-

Dr. Jan Fabio Nickels
-

Dr. Anne Rademacher
-

Sabrina Schumacher
-

Ezgi Sen
-

Simon Steiger
-

Sofie Steinfest
-

Tsz Ching Chloe Tang
-

Claire Vargas
-
Robin Weinmann
-

Sina Jasmin Wille
Selected Publications
Poos AM, Prokoph N, Przybilla MJ, Mallm J-P, Steiger S, Seufert I, John L, Tirier SM, Bauer K, Baumann A, Munawar U, Rasche L, Kortuem M, Giesen N, Huhn S, Reichert P, Mueller-Tidow C, Goldschmidt H, Stegle O, Raab MS, Rippe K, Weinhold N
Trojanowski J, Frank L, Rademacher A, Mücke N, Grigaitis P, Rippe K.
Frank L, Rademacher A, Mücke N, Tirier SM, Koeleman E, Knotz C, Schumacher S, Stainczyk SA, Westermann F, Fröhling S, Chudasama P, Rippe K.
Tirier SM, Mallm J-P, Steiger S, Poos AM, Awwad M, Giesen N, Casiraghi N, Susak H, Bauer K, Baumann A, John L, Seckinger A, Hose D, Müller-Tidow C, Goldschmidt H, Stegle O, Hundemer M, Weinhold N, Raab MS, Rippe K.
Erdel F, Rademacher A, Vlijm R, Tünnermann J, Schweigert E, Frank L, Yserantant K, Hummert J, Bauer C, Schumacher S, Weinmann R, Alwash AA, Normand C, Herten D-P, Engelhardt J, Rippe K.
Get in touch with us

