Pediatric Neurooncology

Prof. Dr. Stefan Pfister
Head of Division
Pediatric Neurooncology is currently a vibrant field of research. This is desperately needed, since brain tumors have become the number one cause of cancer-related mortality in children. Our group aims to bridge the gap between generating genomic screening data as well as faithful models for preclinical drug testing, and exploiting these data for the sake of our patients.
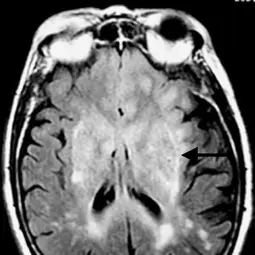
Our research
Pediatric neurooncology - research for better therapies
Brain tumors are the most common cause of cancer-related death in children. While molecular findings have already been successfully integrated into clinical practice in hemato-oncology, neuro-oncology still faces major challenges. Our aim is to better understand the biological basis of childhood brain tumors and to develop new therapeutic approaches.
Our research approach
We analyze tumors genetically using state-of-the-art high-throughput sequencing and make these findings usable for clinical application. This includes the identification and validation of biomarkers that enable predictions to be made about the course of the disease and the response to therapy. We also test new treatment approaches in preclinical models, often in combination with proven chemotherapeutic agents, in order to develop personalized therapies.
Childhood brain tumors are biologically extremely diverse, both between different tumor types and within a single tumor. This diversity makes it difficult to tailor treatments. Our research helps to decipher these genetic and epigenetic differences. Using preclinical models and state-of-the-art sequencing technologies, we investigate tumor development, metastasis and relapse. We are also investigating tumor-specific changes in cerebrospinal fluid and blood plasma in order to develop new diagnostic markers and better monitor therapeutic success.
Our goal: a better understanding of childhood brain tumors and innovative, individual treatment options for affected children.
Team
-

Prof. Dr. Stefan Pfister
Abteilungsleiter
-

Dr. Robert Autry
Gruppenleiter
-

Dr. Kendra Maaß
Gruppenleiterin
-

Dr. Kristian Pajtler
Gruppenleiter
-

Dr. Supat Thongjuea
Gruppenleiter
-

Dr. Marc Zuckermann
Gruppenleiter
Selected publications
Schwalbe EC, Lindsey JC, Danilenko M, Hill RM, Crosier S, Ryan SL, Williamson D, Castle J, Hicks D, Kool M, Milde T, Korshunov A, Pfister SM, Bailey S, Clifford SC.
Sepp M, Leiss K, Murat F, Okonechnikov K, Joshi P, Leushkin E, Spänig L, Mbengue N, Schneider C, Schmidt J, Trost N, Schauer M, Khaitovich P, Lisgo S, Palkovits M, Giere P, Kutscher LM, Anders S, Cardoso-Moreira M, Sarropoulos I, Pfister SM, Kaessmann H.
Ghasemi DR, Okonechnikov K, Rademacher A, Tirier S, Maass KK, …, Jones DTW, Kawauchi D, Fraenkel E, Mallm JP, Rippe K, Korshunov A, Pfister SM, Pajtler KW.
Latest publications
Patient-derived ependymoma and medulloblastoma tumoroids: generation, biobanking and drug screening.
Lago, C. ; Leva, G. ; Kool, M. ; Miele, E. ; Tiberi, L.
Jeong, D. ; Danielli, S. G. ; Maaß, K. K. ; Ghasemi, D. R. ; Tetzlaff, S. ; Reyhan, E. ; Jiang, L. ; Katiyar, S. ; Sundheimer, J. K. ; Lo Cascio, C. ; Neyazi, S. ; de Biagi-Junior, C. A. O. ; Couvillon, E. ; Castellani, S. ; Pazyra-Murphy, M. ; Mullally, M. ; Dehler, M. P. ; Englinger, B. ; Nascimento, A. ; Cruzeiro, G. A. V. ; Marques, J. G. ; Haase, R. D. ; Nguyen, C. M. ; Baumgartner, A.-C. ; Rozowsky, J. S. ; Hack, O. A. ; Shaw, M. L. ; Lotsch-Gojo, D. ; Bruckner, K. ; Korshunov, A. ; Pfister, S. ; Kool, M. ; Nowakowski, T. J. ; Gojo, J. ; Baird, L. ; Alexandrescu, S. ; Pajtler, K. ; Venkataramani, V. ; Filbin, M. G.
Smith, K. S. ; Fischer, T. ; Han, K. ; Kostecka, A. ; Lin, H. ; Senfter, D. ; Soliman, T. ; Stepien, N. ; Volz, S. ; Schwarz, N. ; Wedig, T. ; Madlener, S. ; Haberler, C. ; Dhanda, S. K. ; Upadhyaya, S. A. ; Blackburn, P. R. ; Schmook, M. T. ; de Bont, J. ; Haapasalo, H. ; Dargvainiene, J. ; Leypoldt, F. ; Pfister, S. ; Hulleman, E. ; Orr, B. A. ; Gajjar, A. ; Robinson, G. W. ; Haapasalo, J. ; Nordfors, K. ; Gojo, J. ; Pajtler, K. ; Maass, K. K. ; Northcott, P. A.
Montiel Equihua, C. ; Molenaar, J. J. ; Areso, I. ; Biegel, J. A. ; Blanc, P. ; Chi, S. N. ; Daems, S. ; Danielson, L. ; Drost, J. ; Franke, N. E. ; Frühwald, M. C. ; Gajjar, A. ; Geller, J. I. ; Huang, A. ; Johann, P. D. ; Kearns, P. ; Nysom, K. ; O'Connor, S. ; Ortiz, M. V. ; Parker, J. ; Patel, S. ; Patel, S. ; Roberts, C. W. ; Williamson, D. ; Yi, J. S. ; Pearson, A. D. ; Jenkinson, D. ; Kool, M. ; Bourdeaut, F. ; LifeArc ; Cancer, I. T. f. C. w. ; UK, C. R. ; team, C. G. C. P.
Sarropoulos, I. ; Sepp, M. ; Yamada, T. ; Schäfer, P. S. L. ; Trost, N. ; Schmidt, J. ; Schneider, C. ; Drummer, C. ; Mißbach, S. ; Taskiran, I. I. ; Hecker, N. ; Bravo González-Blas, C. ; Frömel, R. ; Joshi, P. K. ; Leushkin, E. ; Arnskötter, F. ; Leiss, K. ; Okonechnikov, K. ; Lisgo, S. ; Palkovits, M. ; Pääbo, S. ; Cardoso-Moreira, M. ; Kutscher, L. ; Behr, R. ; Pfister, S. ; Aerts, S. ; Kaessmann, H.
Schoonbeek, M. C. ; Gestraud, P. ; Vernooij, L. ; Boltjes, A. ; Amo-Addae, V. ; van Luik, M. ; Del Nery, E. ; Bellini, A. ; Chua, E. ; Swaak, S. ; Looze, E. J. ; Pietiäinen, V. M. ; Turunen, L. L. ; Saarela, J. S. ; Schueler, J. ; Indersie, E. ; Gürgen, D. ; Scotlandi, K. ; Eggert, A. ; Bouarich, R. ; Bourdeaut, F. ; Zaidi, S. ; Surdez, D. ; Carcaboso, Á. M. ; Geoerger, B. ; Iddir, Y. ; Saint-Charles, A. ; Saberi-Ansari, E. ; Cavalli, F. ; Gopisetty, A. ; Rief, E. M. ; Caron, H. N. ; Stancato, L. ; Vassal, G. ; Pfister, S. ; Koster, J. ; Eising, S. ; van Hooff, S. R. ; van den Boogaard, M. L. ; Schleiermacher, G. ; Molenaar, J. J.
Bessler, N. ; Wezenaar, A. K. L. ; Ariese, H. C. R. ; Honhoff, C. ; Dommann, N. ; Wehrens, E. J. ; Ruiz Moreno, C. ; van den Broek, T. J. M. ; Collot, R. V. U. ; Kloosterman, D. J. ; Keramati, F. ; Roosen, M. ; de Blank, S. ; van Vliet, E. ; Barrera Román, M. ; Gatti, L. C. D. E. ; Ertürk, A. ; Kuball, J. ; Sebestyén, Z. ; Kool, M. ; Patrizi, S. ; Miele, E. ; Künkele, A. ; Kranendonk, M. E. G. ; Cornel, A. M. ; Nierkens, S. ; Mayer, C. ; Stunnenberg, H. G. ; Alemany, A. ; Alieva, M. ; Rios, A. C.
Hlevnjak, M. ; Heublein, S. ; Thewes, V. ; Wagener, L. ; Pixberg, C. ; Fremd, C. ; Michel, L. ; Maurer, C. ; Buschhorn, L. ; Dikow, N. ; Feng, F. ; Fröhling, S. ; Herold-Mende, C. ; Hirsch, S. ; Hong, C. ; Hübschmann, D. ; Jassowicz, L. ; Kozyulina, P. ; Pfütze, K. ; Schlenk, R. F. ; Sinn, H.-P. ; Smetanay, K. ; Springfeld, C. ; Stenzinger, A. ; Wagner, C. ; Wolf, S. ; Trumpp, A. ; Jäger, D. ; Zivanovic, O. ; Zapatka, M. ; Schneeweiss, A. ; Lichter, P.
Postlmayr, A. ; Sanchez Bergman, A. ; Torrejon Diaz, J. ; Ciraulo, B. ; Yan, S. ; Hofmann, N. ; Carbajal, S. ; Dobler, R. ; Machaalani, C. ; Schönholzer, M. T. ; Priego Gonzalez, L. ; De Micheli, A. J. ; Berenjeno-Correa, E. ; Baroncini, L. ; Grotzer, M. A. ; Tabori, U. ; Hawkins, C. ; Ayrault, O. ; Zuckermann, M. ; Baumgartner, M. ; Guerreiro Stücklin, A. S.
Kolodziejczak, A. ; Selt, F. ; Peterziel, H. ; Jamaladdin, N. ; Mack, N. ; Maaß, K. ; Meulenbroeks, C. ; Sigaud, R. ; Herold-Mende, C. ; El Damaty, A. ; Burhenne, J. ; Ohmura, S. ; Holland-Letz, T. ; Kutscher, L. M. ; Ernst, A. ; Wei, P.-C. ; Grünewald, T. G. P. ; Oehme, I. ; Kool, M. ; Jones, D. T. W. ; Pajtler, K. W. ; Kratz, C. P. ; Pfister, S. M. ; Witt, O. ; Milde, T.
Collot, R. ; Ruiz-Moreno, C. ; Honhoff, C. ; van den Broek, T. J. M. ; Wezenaar, A. K. L. ; Kloosterman, D. J. ; Ariese, H. C. R. ; Johnson, H. ; Vervoort, B. M. T. ; Jeiroshi, A. ; Bunt, J. ; van Ineveld, R. L. ; Bokobza, E. ; Rebel, H. G. ; Te Pas, B. M. ; Ringnalda, F. C. A. ; van de Wetering, M. ; Robe, P. A. ; Kool, M. ; Cochran, J. R. ; Kranendonk, M. E. G. ; van Vuurden, D. G. ; Hulleman, E. ; Wehrens, E. J. ; Zomer, A. ; Stunnenberg, H. G. ; Rios, A. C.
Gatz, S. A. ; Glade-Bender, J. ; Pearson, A. D. J. ; Ortiz, M. V. ; Bernardi, R. ; Chesler, L. ; Clifford, S. ; Cohen-Gogo, S. ; De La Cuesta, E. ; de Rojas, T. ; Durinck, K. ; Federico, S. ; Fox, E. ; George, S. ; Gounaris, I. ; Henssen, A. G. ; Irwin, M. ; Kool, M. ; Lau, A. ; Nysom, K. ; Pappo, A. ; Pennock, G. K. ; Pfister, S. M. ; Scobie, N. ; Slotkin, E. K. ; Smith, M. ; Speleman, F. ; Stewart, E. A. ; Weigel, B. J. ; Vassal, G.
Fernandez, N. R. ; Chang, Y. ; Nunes, N. M. ; Dimayacyac, J. R. ; Levine, A. ; Ringel, A. ; Negm, L. ; Ercan, A. B. ; Hess, J. M. ; Ahmad, O. ; Lee, C. ; Stengs, L. ; Bianchi, V. ; Edwards, M. ; Doherty, S. ; Chung, J. ; Nobre, L. ; Bennett, J. ; Dodgshun, A. J. ; Jones, D. ; Pfister, S. ; Villani, A. ; Malkin, D. ; Ramaswamy, V. ; Huang, A. ; Bouffet, E. ; Aronson, M. ; Dirks, P. B. ; Shlien, A. ; Getz, G. ; Maruvka, Y. E. ; Ertl-Wagner, B. ; Hawkins, C. ; Das, A. ; Tabori, U.
Funk, C. ; Ehlers, A. ; Orth, M. F. ; Aljakouch, K. ; Li, J. ; Hoelting, T. L. ; Will, R. ; Geyer, F. H. ; Ceranski, A. K. ; Willis, F. ; Vinca, E. ; Ohmura, S. ; Imle, R. ; Siebenlist, J. ; Yershova, A. ; Knott, M. M. ; Zahnow, F. ; Sastre, A. ; Alonso, J. ; Sahm, F. ; Peterziel, H. ; Loboda, A. ; Schneider, M. ; Banito, A. ; Leprivier, G. ; Hartmann, W. ; Dirksen, U. ; Witt, O. ; Oehme, I. ; Pfister, S. M. ; Romero-Pérez, L. ; Krijgsveld, J. ; Cidre-Aranaz, F. ; Grünewald, T. ; Musa, J.
Pfaff, E. ; Schramm, K. ; Blattner-Johnson, M. ; Jones, B. C. ; Stark, S. ; Balasubramanian, G. P. ; Previti, C. ; Autry, R. J. ; Fiesel, P. ; Sahm, F. ; Reuss, D. ; von Deimling, A. ; van Tilburg, C. M. ; Pajtler, K. W. ; Milde, T. ; Dirksen, U. ; Kramm, C. M. ; von Bueren, A. O. ; Hutter, C. ; de Wilde, B. ; Molenaar, J. ; Gerber, N. U. ; Lohi, O. ; Munthe-Kaas, M. C. ; Georgantzi, K. ; Kazanowska, B. ; Zápotocký, M. ; Kattamis, A. ; Filippidou, M. ; Fried, I. ; Pfister, S. M. ; Witt, O. ; Jones, D.
Oak, N. ; Chen, W. ; Blake, A. ; Harrison, L. ; O'Brien, M. ; Previti, C. ; Balasubramanian, G. ; Maass, K. ; Hirsch, S. ; Penkert, J. ; Jones, B. ; Schramm, K. ; Nathrath, M. ; Pajtler, K. ; Jones, D. ; Witt, O. ; Dirksen, U. ; Li, J. ; Sapkota, Y. ; Ness, K. K. ; Guenther, L. M. ; Pfister, S. ; Kratz, C. ; Wang, Z. ; Armstrong, G. T. ; Hudson, M. M. ; Wu, G. ; Autry, R. ; Nichols, K. E. ; Sharma, R.
A spatial map of MAPK-activated immunosuppressive myeloid populations in pediatric low-grade glioma.
Andrade, A. F. ; Sigaud, R. ; Puligandla, E. ; Liu, B. ; Karimi, E. ; Annett, A. ; Rezanejad, M. ; Jawhar, W. ; Taylor, R. ; Cao, Y. ; Schmid, S. ; Selt, F. ; Hernáiz Driever, P. ; Horn, S. ; Sievers, P. ; Prinz, M. ; Glatzel, M. ; Mawrin, C. ; Hartmann, C. ; Monoranu, C.-M. ; Konnikova, L. ; Sahm, F. ; Pfister, S. ; Witt, O. ; Jones, D. ; Koch, A. ; Kleinman, C. L. ; Capper, D. ; Walsh, L. ; Jabado, N. ; Milde, T.
Wick, W. ; Lanz, L.-M. ; Wick, A. ; Harting, I. ; Dettmer, S. ; Suwala, A. K. ; Ketter, R. ; Tabatabai, G. ; Seliger, C. ; Glas, M. ; Burger, M. C. ; Timmer, M. ; Ringel, F. A. ; Mildenberger, I. ; Schulz-Schaeffer, W. J. ; Winkler, F. ; König, L. ; Herold-Mende, C. ; Eisenmenger, A. ; Pfister, S. ; Renovanz, M. ; Bendszus, M. ; Sahm, F. ; Platten, M. ; Kessler, T.
Wasserman, J. D. ; Schneider, K. W. ; Achatz, M.-I. ; Nakano, Y. ; Zelley, K. ; Gallinger, B. ; Bauer, A. J. ; Becktell, K. D. ; Wassner, A. J. ; Raiti, L. ; Doria, A. S. ; States, L. J. ; Stratakis, C. A. ; Brodeur, G. M. ; Diller, L. R. ; Kamihara, J. ; Malkin, D. ; Pajtler, K. ; Tamura, C. ; Villani, A. ; Widjaja, E. ; Das, A. ; Rednam, S. P.
Fischer, T. ; Maass, K. ; Schoof, M. ; Puranachot, P. ; Sievers, P. ; Montigel, S. ; Fluhr, T. ; El Damaty, A. ; Seitz, A. ; Harrabi, S. ; Schramm, K. ; Sturm, D. ; Jones, D. ; Pfister, S. ; Witt, O. ; van Tilburg, C. M. ; Milde, T. ; Schüller, U. ; Pajtler, K.
Blanco-Carmona, E. ; Paassen, I. ; He, J. ; DeMartino, J. ; Büllesbach, A. ; Anderson, N. ; Buhl, J. L. ; Federico, A. ; Mauermann, M. ; Brok, M. ; Straathof, K. ; Behjati, S. ; Vibhakar, R. ; Donson, A. M. ; Foreman, N. K. ; Shaw, M. ; Frühwald, M. C. ; Korshunov, A. ; Hasselblatt, M. ; Thomas, C. ; Franke, N. ; Kranendonk, M. E. ; Hoving, E. W. ; Jäger, N. ; Johann, P. D. ; Pfister, S. ; Filbin, M. G. ; Kool, M. ; Drost, J.
Monnikhof, M. ; Schakelaar, M. Y. ; Meulenbroeks, C. ; Quist, M. ; Perzolli, A. ; Selten, A. ; Koster, C. J. M. ; Maassen, D. S. C. ; Montoro Canelo, A. ; Fredriks, M. ; Koppers, M. J. A. ; Clevers, K. ; Klein, J. ; Kaludjerovic, V. ; Meeldijk, J. ; Pijnappel, E. W. ; Rebel, H. G. ; van Kempen, S. ; Crnko, S. ; Koorman, T. ; Federico, A. ; Valzano, F. ; Wesseling, P. ; Calkoen, F. G. J. ; van der Lugt, J. ; Ten Broeke, T. ; Kool, M. ; Bovenschen, N.
Rubio-San-Simón, A. ; Ritzmann, T. A. ; Obrecht-Sturm, D. ; Benesch, M. ; Timmermann, B. ; Leblond, P. ; Kilday, J.-P. ; Poggi, G. ; Thorp, N. ; Massimino, M. ; van Veelen, M.-L. ; Schuhmann, M. ; Thomale, U.-W. ; Tippelt, S. ; Schüller, U. ; Rutkowski, S. ; Grundy, R. G. ; Bolle, S. ; Fernández-Teijeiro, A. ; Pajtler, K.
Karremann, M. ; Perwein, T. ; von Bueren, A. O. ; Gielen, G. H. ; Benesch, M. ; Nussbaumer, G. ; Friker, L. L. ; Waha, A. ; Sturm, D. ; Jones, D. T. W. ; Witt, O. ; Pfister, S. M. ; Eyrich, M. ; Thomale, U. W. ; Valentini, C. ; Krause, M. ; Van Gool, S. W. ; Frühwald, M. C. ; Hernaiz-Driever, P. ; Ebinger, M. ; Ebetsberger-Dachs, G. ; Crazzolara, R. ; Haberler, C. ; Sahm, F. ; Gerber, N. U. ; Classen, C. F. ; Sträter, R. ; Kerl, K. ; Rutkowski, S. ; Fleischhack, G. ; Buiren, M. v. ; Kortmann, R.-D. ; Hagel, C. ; Calaminus, G. ; Faldum, A. ; Warmuth-Metz, M. ; Bison, B. ; Pietsch, T. ; Hoffmann, M. ; Kwiecien, R. ; Kramm, C. M.
Nozzoli, F. ; Rahmanzade, R. ; Schmid, S. ; Schweizer, L. ; Schrimpf, D. ; Friedel, D. ; Göbel, K. ; Reuss, D. E. ; Banan, R. ; Sievers, P. ; Pusch, S. ; Bogumil, H. ; Hinz, F. ; Suwala, A. K. ; Aras, F. K. ; Friedrich, L. ; Osella-Abate, S. ; Ricci, A. A. ; Macciotta, A. ; Simon, T. ; Fleischhack, G. ; Keyvani, K. ; Hansford, J. R. ; Khuong-Quang, D.-A. ; Schucht, P. ; Maragkou, T. ; Juratli, T. A. ; Meinhardt, M. ; Zechel, S. ; Stadelmann, C. ; Coras, R. ; Sakowitz, O. W. ; Goeppert, B. ; Schittenhelm, J. ; Etminan, N. ; Ratliff, M. ; Herold-Mende, C. ; Pfister, S. ; Wick, W. ; Krieg, S. M. ; von Deimling, A. ; Sahm, F. ; Bertero, L.
Hemminki, K. ; Li, X. ; Sundquist, K. ; Sundquist, J. ; Hirsch, S. ; Ji, J. ; Hemminki, A. ; Försti, A.
Fischer, T. ; Maass, K. ; Puranachot, P. ; Mieskolainen, M. ; Sill, M. ; Schad, P. ; Volz, S. ; Rosing, F. ; Wedig, T. ; Schwarz, N. ; Finster, A. ; Iser, F. ; Meyer, J. ; Sahm, F. ; Lohi, O. ; Damaty, A. E. ; Brors, B. ; Haapasalo, H. ; Pfister, S. ; Haapasalo, J. ; Pajtler, K. ; Nordfors, K.
Talbot, J. ; Fombonne, J. ; Torrejon, J. ; Babcock, B. R. ; McSwain, L. F. ; Rama, N. ; Lospinoso Severini, L. ; Bonerandi, E. ; Marsaud, V. ; Bernardi, F. ; Gharsalli, T. ; Guix, C. ; Ducarouge, B. ; Neururer, V. ; Basili, I. ; Mercier, A. L. ; Yu, H. ; Forget, A. ; Indersie, E. ; Leboucher, S. ; Souphron, J. ; Okonechnikov, K. ; Wang, W. ; Kawauchi, D. ; Wainwright, B. J. ; Frappaz, D. ; Varlet, P. ; Dufour, C. ; Beccaria, K. ; Blauwblomme, T. ; Martignetti, L. ; Di Marcotullio, L. ; Puget, S. ; Doz, F. ; Bourdeaut, F. ; Masliah-Planchon, J. ; Gershon, T. R. ; Mehlen, P. ; Ayrault, O.
Deng, M. Y. ; Maas, S. L. N. ; Anil, G. ; Sievers, P. ; Lischalk, J. ; Zhao, E. ; Rauh, S. ; Jessen, I. ; Eichkorn, T. ; Regnery, S. ; Bauer, L. ; Held, T. ; Hoegen-Sassmannshausen, P. ; Seidensaal, K. ; Hörner-Rieber, J. ; Pfister, S. M. ; Wick, A. ; Wick, W. ; von Deimling, A. ; Herfarth, K. ; Jungk, C. ; Krieg, S. M. ; Debus, J. ; Sahm, F. ; König, L.
Beck, A. ; Gabler-Pamer, L. ; Alencastro Veiga Cruzeiro, G. ; Lambo, S. ; Englinger, B. ; Shaw, M. L. ; Hack, O. A. ; Liu, I. ; Haase, R. D. ; de Biagi, C. A. O. ; Baumgartner, A. ; Nascimento Silva, A. D. ; Klenner, M. ; Freidel, P. S. ; Herms, J. ; von Baumgarten, L. ; Tonn, J. C. ; Thon, N. ; Bruckner, K. ; Madlener, S. ; Mayr, L. ; Senfter, D. ; Peyrl, A. ; Slavc, I. ; Lötsch, D. ; Dorfer, C. ; Geyregger, R. ; Amberg, N. ; Haberler, C. ; Mack, N. ; Schwalm, B. ; Pfister, S. ; Korshunov, A. ; Baird, L. C. ; Yang, E. ; Chi, S. N. ; Alexandrescu, S. ; Gojo, J. ; Kool, M. ; Hovestadt, V. ; Filbin, M. G.
Ahmad, S. T. ; Li, Y. ; Garcia-Lopez, J. ; Gudenas, B. L. ; Hadley, J. ; Paul, L. ; Wu, S. C. ; Refaat, A. ; Kojic, M. ; Batts, M. ; Soliman, T. ; Pitre, A. ; Arnskötter, F. ; Zindy, F. ; Jones, A. ; Twarog, N. R. ; Mayasundari, A. ; Bianski, B. ; Tinkle, C. ; Shirinifard, A. ; Janke, L. ; Lu, M. ; Lewis, S. A. ; Onar-Thomas, A. ; Pfister, S. M. ; Gajjar, A. ; Baker, S. J. ; Roussel, M. F. ; Rankovic, Z. ; Robinson, G. W. ; Orr, B. A. ; Wainwright, B. ; Shelat, A. A. ; Waszak, S. M. ; Kutscher, L. M. ; Lin, H. ; Northcott, P. A.
Okonechnikov, K. ; Joshi, P. K. ; Körber, V. ; Rademacher, A. ; Bortolomeazzi, M. ; Mallm, J.-P. ; Vaillant, J. ; da Silva, P. B. G. ; Statz, B. ; Sepp, M. ; Sarropoulos, I. ; Yamada, T. ; Wittmann, A. ; Schramm, K. ; Blattner-Johnson, M. ; Fiesel, P. ; Jones, B. ; Jäger, N. ; Milde, T. ; Pajtler, K. ; van Tilburg, C. M. ; Witt, O. ; Bochennek, K. ; Weber, K. J. ; Nonnenmacher, L. ; Reimann, C. ; Ghasemi, D. R. ; Schüller, U. ; Mynarek, M. ; Rutkowski, S. ; Jones, D. ; Korshunov, A. ; Rippe, K. ; Westermann, F. ; Thongjuea, S. ; Höfer, T. ; Kaessmann, H. ; Kutscher, L. ; Pfister, S.
LaBelle, J. J. ; Haase, R. D. ; Beck, A. ; Haase, J. ; Jiang, L. ; Oliveira de Biagi, C. A. ; Neyazi, S. ; Englinger, B. ; Liu, I. ; Trissal, M. ; Jeong, D. ; Hack, O. A. ; Nascimento, A. ; Shaw, M. L. ; Nguyen, C. M. ; Castellani, S. ; Mathewson, N. D. ; Ashenberg, O. ; Veiga Cruzeiro, G. A. ; Rosenberg, T. ; Vogelzang, J. R. ; Pyrdol, J. ; Marx, S. ; Luomo, A. M. ; Godicelj, A. ; Baumgartner, A. ; Rozowsky, J. S. ; Madlener, S. ; Mayr, L. ; Peyrl, A. ; Geyeregger, R. ; Loetsch, D. ; Dorfer, C. ; Haberler, C. ; Stepien, N. ; Slavc, I. ; Davidson, T. B. ; Prins, R. M. ; Yeo, K. K. ; Cooney, T. ; Ligon, K. ; Lidov, H. ; Alexandrescu, S. ; Baird, L. C. ; Gojo, J. ; Wucherpfennig, K. W. ; Filbin, M. G.
Rubio-San-Simón, A. ; Ritzmann, T. A. ; Obrecht-Sturm, D. ; Benesch, M. ; Timmermann, B. ; Leblond, P. ; Kilday, J.-P. ; Poggi, G. ; Thorp, N. ; Massimino, M. ; van Veelen, M.-L. ; Schuhmann, M. ; Thomale, U.-W. ; Tippelt, S. ; Schüller, U. ; Rutkowski, S. ; Grundy, R. G. ; Bolle, S. ; Fernández-Teijeiro, A. ; Pajtler, K.
Park, S. H. ; Sugier, P.-E. ; Asgari, Y. ; Karimi, M. ; Kaaks, R. ; Fortner, R. T. ; Schulze, M. ; Agnoli, C. ; Pasanisi, F. ; Sacerdote, C. ; Rodriguez-Barranco, M. ; Aizpurua, A. ; Cabrera Castro, N. ; Guevara, M. ; Tin Tin, S. ; Weiderpass, E. ; de Vathaire, F. ; Lesueur, F. ; Guénel, P. ; Mulot, C. ; Laurent-Puig, P. ; Ostroumova, E. ; Boland-Auge, A. ; Deleuze, J.-F. ; Thomsen, H. ; Försti, A. ; Elisei, R. ; Gemignani, F. ; Landi, S. ; Rinaldi, S. ; Elbaz, A. ; Domenighetti, C. ; Truong, T.
Geyer, F. ; Ritter, A. ; Kinn-Gurzo, S. ; Faehling, T. ; Li, J. ; Jarosch, A. ; Ngo, C. ; Vinca, E. ; Aljakouch, K. ; Orynbek, A. ; Ohmura, S. ; Kirchner, T. ; Imle, R. ; Romero-Pérez, L. ; Díaz-Martín, J. ; Bertram, S. ; de Álava, E. ; Henon, C. ; Postel-Vilnay, S. ; Banito, A. ; Sill, M. ; Versleijen-Jonkers, Y. ; Mayer, B. F. B. ; Ebinger, M. ; Sparber-Sauer, M. ; Stegmaier, S. ; Baumhoer, D. ; Hartmann, W. ; Krijgsveld, J. ; Horst, D. ; Delattre, O. ; Grohar, P. J. ; Grünewald, T. ; Cidre-Aranaz, F.
Hirsch, S. ; Rahmanzade, R. ; Grund, K. ; Sutter, C. ; Schramm, K. ; Selt, F. ; Ecker, J. ; Jones, B. C. ; Schrimpf, D. ; Demmert, M. ; Guerreiro Stücklin, A. S. ; Hernaiz Driever, P. ; Mezger, M. ; Brecht, I. ; Adib, S. D. ; Brummel, B. ; Sturm, D. ; Dikow, N. ; Hempel, M. ; Milde, T. ; Pajtler, K. ; Jones, D. T. W. ; Pfister, S. M. ; von Deimling, A. ; Sahm, F. ; Schaaf, C. P.
Mateos, M. K. ; Ajuyah, P. ; Fuentes-Bolanos, N. ; El-Kamand, S. ; Barahona, P. ; Altekoester, A.-K. ; Mayoh, C. ; Holliday, H. ; Liu, J. ; Cui, L. ; Pfaff, E. ; Mackay, A. ; Resnick, A. C. ; Pinese, M. ; Lau, L. M. S. ; Khuong-Quang, D.-A. ; Dias, K. ; Goudie, C. ; Salkeld, A. ; Rokita, J. L. ; Jones, D. T. W. ; Juretic, N. ; Hayden, E. ; Pfister, S. M. ; Kramm, C. M. ; Blattner-Johnson, M. ; Jabado, N. ; Tsoli, M. ; Vittorio, O. ; Mueller, S. ; Guo, Y. ; Tucker, K. ; Waszak, S. M. ; Perreault, S. ; Jones, C. ; Wong-Erasmus, M. ; Cowley, M. J. ; Ziegler, D. S.
Delaidelli, A. ; Burwag, F. ; Ben-Neriah, S. ; Suk, Y. ; Shyp, T. ; Kosteniuk, S. ; Dunham, C. ; Cheng, S. ; Okonechnikov, K. ; Schrimpf, D. ; von Deimling, A. ; Ellezam, B. ; Perreault, S. ; Singh, S. ; Hawkins, C. ; Kool, M. ; Pfister, S. ; Steidl, C. ; Hughes, C. ; Korshunov, A. ; Sorensen, P. H.
Salman, Z. ; Moreira, D. C. ; Ul Ain, R. ; Hoveyan, J. ; Resendiz, A. E. B. ; Rahmartani, L. D. ; Zhang, A. ; Amayiri, N. ; Bailey, S. ; Bouffet, E. ; Chan, G. C. ; Liu, A. P. ; La Madrid, A. M. ; Mushtaq, N. ; Tsui, K. ; Ngcana, T. V. Z. ; Salazar, M. S. ; Bhat K, V. ; Cirt, R. ; Somathilaka, M. ; Yang, P. ; Chinnaswamy, G. ; Dhall, G. ; Gupta, T. ; Jalali, R. ; Lassaletta, A. ; Osorio, D. S. ; Shatara, M. ; Upadhyaya, S. A. ; Uppuluri, R. ; Pfister, S. ; Ybarra, S. ; DiNovis, E. ; Rodriguez-Galindo, C. ; Qaddoumi, I.
Scheck, J. ; Boztepe, B. ; Kernbach, J. M. ; Karimian-Jazi, K. ; Heinz, L. ; Schregel, K. ; Sturm, V. ; Schell, M. ; Bojcevski, J. ; Fischer, M. ; Eurich, R. ; Poschke, I. ; Schwarz, J. ; Agardy, D. A. ; Jünger, S. ; Schulz, C. ; Althammer, F. ; Osidach, A. R. ; Abdollahi, A. ; Bunse, L. ; Venkataramani, V. ; Pfister, S. M. ; Winkler, F. ; Kessler, T. ; Wick, W. ; Heiland, S. ; Platten, M. ; Rodell, C. ; Bendszus, M. ; Weidenfeld, I. ; Breckwoldt, M.
Taylor, A. M. ; Lombardi, J. T. ; Patel, A. J. ; Tamariz, A. ; Martin, J. ; Bookland, M. J. ; Hersh, D. S. ; Cantor, E. ; Song, X. ; Sahm, F. ; Ng, P. K. ; Gell, J. J. ; Lau, C. C.
Hinz, F. ; Friedel, D. ; Ippen, F. ; Sill, M. ; Korshunov, A. ; Schweizer, L. ; Schrimpf, D. ; Göbel, K. ; Friedrich, L. S. ; Aras, F. K. ; Bogumil, H. ; Banan, R. ; Dohmen, H. ; Acker, T. ; Brandner, S. ; Schmid, S. ; Capper, D. ; Grassl, N. ; Boldt, H. B. ; Wesseling, P. ; Maas, S. L. N. ; Martinez, J. P. G. ; Stadelmann, C. ; Reifenberger, G. ; Stehle, T. ; Barrantes-Freer, A. ; Juratli, T. A. ; Pusch, S. ; Haag, D. ; Reuss, D. E. ; Herold-Mende, C. ; Krieg, S. ; Wick, W. ; Etminan, N. ; Platten, M. ; Pfister, S. M. ; Jones, D. T. W. ; Sahm, F. ; von Deimling, A. ; Suwala, A. K.
Gerstung, M. ; Jin, D. ; Shmatko, A. ; Patel, A. ; Rahmanzade, R. ; Banan, R. ; Friedrich, L. ; Sievers, P. ; Hamelmann, S. ; Schrimpf, D. ; Göbel, K. ; Bogumil, H. ; Maas, S. ; Sill, M. ; Hinz, F. ; Suwala, A. ; Keller, F. ; Habel, A. ; Rukhovich, G. ; Rutz, S. ; Alhalabi, O. ; Sturm, D. ; Reuss, D. E. ; Wesseling, P. ; Woehrer, A. ; Heppner, F. ; Herold-Mende, C. ; Krieg, S. ; Wick, W. ; Jones, D. ; Pfister, S. ; Al-Hussaini, M. ; Hou, Y. ; Costa, F. D. ; Ille, S. ; Sehrig, J. ; Piovesan-Lago, P. ; Suchorska, B. ; Ahmad, O. ; Schweizer, L. ; Bertero, L. ; Acker, T. ; Tauziède-Espariat, A. ; Varlet, P. ; Brandner, S. ; Bai, X. ; von Deimling, A. ; Sahm, F.
Fürst, A. ; Ruf, V. ; Fiedler, C. ; Rutkowski, S. ; Sill, M. ; Korshunov, A. ; Gerber, N. U. ; Frank, S. ; Hench, J. ; Schüller, U.
Chavaz, L. ; Bagchi, A. ; Dhanda, S. K. ; Toutain, F. ; Pfister, S. M. ; Sturm, D. ; Pietsch, T. ; Gielen, G. H. ; Waha, A. ; Clarke, M. ; Lu, C. ; Karremann, M. ; Benesch, M. ; Perwein, T. ; Nussbaumer, G. ; Kramm, C. ; Massimino, M. ; Biassoni, V. ; Vinci, M. ; Mastronuzzi, A. ; van Vuurden, D. ; Veldhuijzen van Zanten, S. E. M. ; Mackay, A. ; Jones, C. ; Jones, D. T. W. ; Guerreiro Stucklin, A. S. ; Tabori, U. ; Hawkins, C. ; Ryall, S. ; Morales La Madrid, A. ; Lassaletta, A. ; Bailey, S. ; Hargrave, D. ; Chiang, J. ; El-Ayadi, M. ; Mançano, B. M. ; Manuel Reis, R. ; Hagel, C. ; Gorsi, H. ; Silvestrini, N. ; Gilani, A. ; Papusha, L. ; Klimo, P. ; Zhou, X. ; Gajjar, A. ; Robinson, G. W. ; von Bueren, A. O.
Valentini, C. ; Perwein, T. ; Bison, B. ; Gielen, G. H. ; Knerlich-Lukoschus, F. ; Bock, H. C. ; Seidel, C. ; Kortmann, R. D. ; Sturm, D. ; Benesch, M. ; Nussbaumer, G. ; Krischer, J. M. ; V Bueren, A. ; Eyrich, M. ; Friker, L. L. ; Hoffmann, M. ; Gkika, E. ; Wittig-Sauerwein, A. ; Hörner-Rieber, J. ; Schwarz, R. ; Jablonska, K. ; Hoffmann, W. ; Vordermark, D. ; Rieken, S. ; Höng, L. ; Rödel, C. ; Timmermann, B. ; Fennell, J. T. ; Claviez, A. ; Karremann, M. ; Kramm, C. M. ; Krause, M.
Wang, Y. ; Liu, K. ; Zhang, Y. ; Novak, D. ; Federico, A. ; Xu, C. ; Horschitz, S. ; Vierthaler, M. ; Sun, Q. ; Wang, N. ; Poelchen, J. ; Steinfass, T. ; Hüser, L. ; Mall, M. ; Umansky, V. ; Utikal, J.
Imle, R. ; Blösel, D. ; Kommoss, F. K. ; Placke, S. ; Stutheit-Zhao, E. ; Blume, C. ; Lupar, D. ; Schmitt, L. ; Winter, C. ; Wagner, L. ; von Eicke, M. ; Walzer, H. ; Förderer, J. ; Laier, S. ; Hertwig, M. ; Peterziel, H. ; Oehme, I. ; Scheuerman, S. ; Seitz, C. ; Geyer, F. H. ; Cidre-Aranaz, F. ; Grünewald, T. ; Vokuhl, C. ; Chudasama, P. ; Scholl, C. ; Schmidt, C. ; Günther, P. ; Sill, M. ; Jones, K. B. ; Pfister, S. ; Autry, R. ; Banito, A.
Hawkins, C. ; Aldape, K. ; Capper, D. ; von Deimling, A. ; Giannini, C. ; Gilbert, M. R. ; Jacques, T. S. ; Jones, D. ; Komori, T. ; Louis, D. N. ; Mueller, S. ; Nasrallah, M. ; Orr, B. A. ; Perry, A. ; Pfister, S. M. ; Sahm, F. ; Sarkar, C. ; Snuderl, M. ; Solomon, D. ; Varlet, P. ; Wesseling, P. ; Reifenberger, G.
Morfouace, M. ; Bielle, F. ; Razis, E. ; Estrade, F. ; Rubio, A. ; Bautista, F. ; de Rojas, T. ; Vieito, M. ; Meade, S. ; Sanson, M. ; Marques, A. C. ; Preusser, M. ; Hatcher, H. ; Balasubramanian, G. P. ; Pineda, E. ; D'Hondt, L. ; Duerinck, J. ; Michotte, A. ; Mawrin, C. ; Ribalta, T. ; Marucci, G. ; Golfinopoulos, V. ; Pfister, S. M. ; Jones, D. T. ; McCabe, M. G.
Sievers, P. ; Arora, S. ; Hielscher, T. ; Savran, D. ; Schrimpf, D. ; Banan, R. ; Vonhören, D. ; Pusch, S. ; Sill, M. ; Appay, R. ; Wirsching, H.-G. ; Hortobagyi, T. ; Dohmen, H. ; Acker, T. ; Kohlhof-Meinecke, P. ; Schweizer, L. ; Wefers, A. K. ; Harter, P. ; Hartmann, C. ; Beschorner, R. ; Schittenhelm, J. ; Behling, F. ; Tabatabai, G. ; Mawrin, C. ; Snuderl, M. ; Maas, S. L. N. ; Wesseling, P. ; Brandner, S. ; Korshunov, A. ; Ratliff, M. ; Krieg, S. M. ; Wick, W. ; Jones, D. T. W. ; Pfister, S. M. ; Holland, E. C. ; von Deimling, A. ; Szulzewsky, F. ; Sahm, F.
Achatz, M. I. ; Villani, A. ; Bertuch, A. A. ; Bougeard, G. ; Chang, V. Y. ; Doria, A. S. ; Gallinger, B. ; Godley, L. A. ; Greer, M. C. ; Kamihara, J. ; Khincha, P. P. ; Kohlmann, W. K. ; Kratz, C. P. ; MacFarland, S. P. ; Maese, L. D. ; Maxwell, K. N. ; Mitchell, S. G. ; Nakano, Y. ; Pfister, S. ; Wasserman, J. D. ; Woodward, E. R. ; Garber, J. E. ; Malkin, D.
Perrino, M. R. ; Jongmans, M. C. J. ; Tomlinson, G. E. ; Greer, M. C. ; Scollon, S. R. ; Mitchell, S. G. ; Hansford, J. R. ; Schultz, K. A. P. ; Kohlmann, W. K. ; Kalish, J. M. ; MacFarland, S. P. ; Das, A. ; Maxwell, K. N. ; Pfister, S. ; Weksberg, R. ; Michaeli, O. ; Tabori, U. ; Ney, G. M. ; Lupo, P. J. ; Brzezinski, J. J. ; Stewart, D. R. ; Woodward, E. R. ; Kratz, C. P.
Velz, J. ; Freudenmann, L. ; Medici, G. ; Dubbelaar, M. ; Mohme, M. ; Ghasemi, D. R. ; Scheid, J. ; Kowalewski, D. J. ; Patterson, A. B. ; Zeitlberger, A. M. ; Lamszus, K. ; Westphal, M. ; Eyrich, M. ; Messing-Jünger, M. ; Röhrig, A. ; Reinhard, H. ; Beccaria, K. ; Craveiro, R. B. ; Frey, B. M. ; Sill, M. ; Nahnsen, S. ; Gauder, M. ; Kapolou, K. ; Silginer, M. ; Weiss, T. ; Wirsching, H.-G. ; Roth, P. ; Grotzer, M. ; Krayenbühl, N. ; Bozinov, O. ; Regli, L. ; Rammensee, H.-G. ; Rushing, E. J. ; Sahm, F. ; Walz, J. ; Weller, M. ; Neidert, M. C.
Aldape, K. ; Capper, D. ; von Deimling, A. ; Giannini, C. ; Gilbert, M. R. ; Hawkins, C. ; Hench, J. ; Jacques, T. S. ; Jones, D. ; Louis, D. N. ; Mueller, S. ; Orr, B. A. ; Nasrallah, M. ; Pfister, S. ; Sahm, F. ; Snuderl, M. ; Solomon, D. ; Varlet, P. ; Wesseling, P.
Hinz, F. ; Friedel, D. ; Korshunov, A. ; Ippen, F. ; Bogumil, H. ; Banan, R. ; Brandner, S. ; Hasselblatt, M. ; Boldt, H. B. ; Dirse, V. ; Dohmen, H. ; Aronica, E. ; Brodhun, M. ; Broekman, M. L. D. ; Capper, D. ; Cherkezov, A. ; Deng, M. Y. ; van Dis, V. ; Felsberg, J. ; Frank, S. ; French, P. J. ; Gerlach, R. ; Göbel, K. ; Goold, E. ; Hench, J. ; Kantelhardt, S. ; Kohlhof-Meinecke, P. ; Krieg, S. ; Mawrin, C. ; Morrison, G. ; Mühlebner, A. ; Ozduman, K. ; Pfister, S. ; Poliani, P. L. ; Prinz, M. ; Reifenberger, G. ; Riemenschneider, M. J. ; Sankowski, R. ; Schrimpf, D. ; Sill, M. ; Snuderl, M. ; Verdijk, R. M. ; Voisin, M. R. ; Wesseling, P. ; Wick, W. ; Reuss, D. E. ; von Deimling, A. ; Sahm, F. ; Maas, S. L. N. ; Suwala, A. K.
Ritter, M. ; Blume, C. ; Tang, Y. ; Patel, A. J. ; Patel, B. ; Berghaus, N. ; Kada Benotmane, J. ; Kueckelhaus, J. ; Yabo, Y. ; Zhang, J. ; Grabis, E. ; Villa, G. ; Zimmer, D. N. ; Khriesh, A. ; Sievers, P. ; Seferbekova, Z. ; Hinz, F. ; Ravi, V. M. ; Seiz-Rosenhagen, M. ; Ratliff, M. ; Herold-Mende, C. ; Schnell, O. ; Beck, J. ; Wick, W. ; von Deimling, A. ; Gerstung, M. ; Heiland, D. H. ; Sahm, F.
Smirnov, P. ; Przybilla, M. J. ; Simovic-Lorenz, M. ; Parra, R. G. ; Susak, H. ; Aichmüller-Ratnaparkhe, M. ; Wong, J. K. ; Körber, V. ; Mallm, J.-P. ; Philippos, G. ; Sill, M. ; Kolb, T. ; Kumar, R. ; Casiraghi, N. ; Okonechnikov, K. ; Ghasemi, D. R. ; Maaß, K. K. ; Pajtler, K. ; Jauch, A. ; Korshunov, A. ; Höfer, T. ; Zapatka, M. ; Pfister, S. ; Huber, W. ; Stegle, O. ; Ernst, A.
Venkataramani, V. ; Yang, Y. ; Ille, S. ; Suchorska, B. ; Loges, S. ; Tost, H. ; Sahm, F. ; Pfister, S. ; Trumpp, A. ; Krieg, S. M. ; Kuner, T. ; Wick, W. ; Winkler, F.
Pfaff, E. ; Schramm, K. ; Blattner-Johnson, M. ; Jones, B. C. ; Stark, S. ; Balasubramanian, G. P. ; Previti, C. ; Autry, R. J. ; Fiesel, P. ; Sahm, F. ; Reuss, D. ; von Deimling, A. ; van Tilburg, C. M. ; Pajtler, K. W. ; Milde, T. ; Dirksen, U. ; Kramm, C. M. ; von Bueren, A. O. ; Munthe-Kaas, M. C. ; Øra, I. ; Pfister, S. M. ; Witt, O. ; Jones, D.
Laemmerer, A. ; Lehmann, C. ; Mayr, L. ; Bruckner, K. ; Gabler, L. ; Senfter, D. ; Meyer, P. ; Balber, T. ; Pirker, C. ; Jaunecker, C. N. ; Kirchhofer, D. ; Vician, P. ; Griesser, M. ; Spiegl-Kreinecker, S. ; Schmook, M. T. ; Traub-Weidinger, T. ; Kuess, P. ; Eckert, F. ; Federico, A. ; Madlener, S. ; Stepien, N. ; Robl, B. ; Baumgartner, A. ; Hainfellner, J. A. ; Dieckmann, K. ; Dorfer, C. ; Roessler, K. ; Corsini, N. S. ; Holzmann, K. ; Schmidt, W. M. ; Peyrl, A. ; Azizi, A. A. ; Haberler, C. ; Beck, A. ; Pfister, S. M. ; Schueler, J. ; Loetsch-Gojo, D. ; Knoblich, J. A. ; Berger, W. ; Gojo, J.
Brzezinski, J. J. ; Becktell, K. D. ; Bougeard, G. ; Brodeur, G. M. ; Diller, L. R. ; Doria, A. S. ; Hansford, J. R. ; Kohlmann, W. K. ; Kratz, C. P. ; MacFarland, S. P. ; Pajtler, K. ; Rednam, S. P. ; Schienda, J. ; States, L. J. ; Villani, A. ; Weksberg, R. ; Zelley, K. ; Tomlinson, G. E. ; Kalish, J. M.
Lim-Fat, M. J. ; Bennett, J. ; Ostrom, Q. ; Touat, M. ; Franceschi, E. ; Schulte, J. ; Bindra, R. S. ; Fangusaro, J. ; Dhall, G. ; Nicholson, J. ; Jackson, S. ; Davidson, T. B. ; Calaminus, G. ; Robinson, G. ; Whittle, J. R. ; Hau, P. ; Ramaswamy, V. ; Pajtler, K. ; Rudà, R. ; Foreman, N. K. ; Hervey-Jumper, S. L. ; Das, S. ; Dirks, P. ; Bi, W. L. ; Huang, A. ; Merchant, T. E. ; Fouladi, M. ; Aldape, K. ; Van den Bent, M. J. ; Packer, R. J. ; Miller, J. J. ; Reardon, D. A. ; Chang, S. M. ; Haas-Kogan, D. ; Tabori, U. ; Hawkins, C. ; Monje, M. ; Wen, P. Y. ; Bouffet, E. ; Yeo, K. K.
Engertsberger, L. ; Benesch, M. ; Mynarek, M. ; Tonn, S. ; Obrecht-Sturm, D. ; Perwein, T. ; Stickan-Verfürth, M. ; Funk, A. ; Timmermann, B. ; Bockmayr, M. ; Eckhardt, A. ; Claviez, A. ; Kortmann, R.-D. ; Riemenschneider, M. J. ; Pietsch, T. ; Bison, B. ; Warmuth-Metz, M. ; Pajtler, K. W. ; Rutkowski, S. ; Schüller, U.
Joshi, P. K. ; Stelzer, T. ; Okonechnikov, K. ; Sarropoulos, I. ; Sepp, M. ; Pour-Jamnani, M. V. ; Rademacher, A. ; Yamada-Saito, T. ; Schneider, C. ; Schmidt, J. ; Schäfer, P. ; Leiss, K. ; Bortolomeazzi, M. ; Mallm, J.-P. ; da Silva, P. B. G. ; Statz, B. ; Wittmann, A. ; Schramm, K. ; Blattner-Johnson, M. ; Fiesel, P. ; Jones, B. ; Milde, T. ; Pajtler, K. ; van Tilburg, C. M. ; Witt, O. ; Rippe, K. ; Korshunov, A. ; Jones, D. T. W. ; Hovestadt, V. ; Northcott, P. A. ; Thongjuea, S. ; Jäger, N. ; Kaessmann, H. ; Pfister, S. M. ; Kutscher, L.
Okonechnikov, K. ; Joshi, P. K. ; Körber, V. ; Rademacher, A. ; Bortolomeazzi, M. ; Mallm, J.-P. ; da Silva, P. B. G. ; Statz, B. ; Sepp, M. ; Sarropoulos, I. ; Yamada-Saito, T. ; Vaillant, J. ; Wittmann, A. ; Schramm, K. ; Blattner-Johnson, M. ; Fiesel, P. ; Jones, B. ; Milde, T. ; Pajtler, K. ; van Tilburg, C. M. ; Witt, O. ; Bochennek, K. ; Weber, K. J. ; Nonnenmacher, L. ; Reimann, C. ; Schüller, U. ; Mynarek, M. ; Rutkowski, S. ; Jones, D. T. W. ; Korshunov, A. ; Rippe, K. ; Westermann, F. ; Thongjuea, S. ; Höfer, T. ; Kaessmann, H. ; Kutscher, L. ; Pfister, S.
Kats, I. ; Simovic-Lorenz, M. ; Schreiber, H. ; Sant, P. ; Mallm, J.-P. ; Körber, V. ; Li, A. ; Velmurugan, P. ; Heuer, S. ; Kües, L. ; Devens, F. ; Sill, M. ; Jugold, M. ; Moustafa, M. ; Abdollahi, A. ; Winkler, F. ; Korshunov, A. ; Pfister, S. ; Stegle, O. ; Ernst, A.
Smirnov, P. ; Przybilla, M. J. ; Simovic-Lorenz, M. ; Parra, R. G. ; Susak, H. ; Aichmüller-Ratnaparkhe, M. ; Wong, J. K. ; Körber, V. ; Mallm, J.-P. ; Philippos, G. ; Sill, M. ; Kolb, T. ; Kumar, R. ; Casiraghi, N. ; Okonechnikov, K. ; Ghasemi, D. R. ; Maaß, K. K. ; Pajtler, K. ; Jauch, A. ; Korshunov, A. ; Höfer, T. ; Zapatka, M. ; Pfister, S. ; Huber, W. ; Stegle, O. ; Ernst, A.
Tauziède-Espariat, A. ; Friker, L. L. ; Nussbaumer, G. ; Bison, B. ; Dangouloff-Ros, V. ; Métais, A. ; Sumerauer, D. ; Zamecnik, J. ; Benesch, M. ; Perwein, T. ; van Vuurden, D. ; Wesseling, P. ; La Madrid, A. M. ; Garrè, M. L. ; Antonelli, M. ; Giangaspero, F. ; Pietsch, T. ; Sturm, D. ; Jones, D. T. W. ; Pfister, S. M. ; Grabovska, Y. ; Mackay, A. ; Jones, C. ; Grill, J. ; Ajlil, Y. ; von Bueren, A. O. ; Karremann, M. ; Hoffmann, M. ; Kramm, C. M. ; Kwiecien, R. ; Castel, D. ; Gielen, G. H. ; Varlet, P.
Perrino, M. R. ; Das, A. ; Scollon, S. R. ; Mitchell, S. G. ; Greer, M. C. ; Yohe, M. E. ; Hansford, J. R. ; Kalish, J. M. ; Schultz, K. A. P. ; MacFarland, S. P. ; Kohlmann, W. K. ; Lupo, P. J. ; Maxwell, K. N. ; Pfister, S. ; Weksberg, R. ; Michaeli, O. ; Jongmans, M. C. J. ; Tomlinson, G. E. ; Brzezinski, J. ; Tabori, U. ; Ney, G. M. ; Gripp, K. W. ; Gross, A. M. ; Widemann, B. C. ; Stewart, D. R. ; Woodward, E. R. ; Kratz, C. P.
Filippidou, M. ; Glentis, S. ; Binenbaum, I. ; Sill, M. ; Roka, K. ; Vlachou, A. ; Avgerinou, G. ; Ecker, J. ; Selt, F. ; Hasselblatt, M. ; Blattner-Johnson, M. ; Schramm, K. ; Trougkou, C. ; Doganis, D. ; Katzilakis, N. ; Ridola, V. ; Papakonstantinou, E. ; Papadakis, V. ; Hatzipantelis, E. ; Kokkinou, E. ; Pons, R. ; Kanaka-Gantenbein, C. ; Sturm, D. ; Hirsch, S. ; Dikow, N. ; Pajtler, K. ; van Tilburg, C. M. ; Frühwald, M. C. ; Milde, T. ; Witt, O. ; Jones, D. ; von Deimling, A. ; Sahm, F. ; Stefanaki, K. ; Pfister, S. ; Kattamis, A.
Fincke, V. E. ; Steinbügl, M. ; Chun, H. E. ; Nemes, K. ; Mucha, M. ; Loßner, M. ; Dorn, F. ; Gastberger, K. ; Bühner, S. ; Sill, M. ; Kröncke, T. ; Siebert, R. ; Melchior, P. ; Furtwängler, R. ; Schlesner, M. ; Vokuhl, C. ; Röcken, C. ; Johann, P. ; Frühwald, M. C.
Ghasemi, D. R. ; Okonechnikov, K. ; Rademacher, A. ; Tirier, S. ; Maass, K. ; Schumacher, H. ; Joshi, P. K. ; Gold, M. P. ; Sundheimer, J. ; Statz, B. ; Rifaioglu, A. S. ; Bauer, K. ; Schumacher, S. ; Bortolomeazzi, M. ; Giangaspero, F. ; Ernst, K. ; Clifford, S. C. ; Saez-Rodriguez, J. ; Jones, D. ; Kawauchi, D. ; Fraenkel, E. ; Mallm, J.-P. ; Rippe, K. ; Korshunov, A. ; Pfister, S. ; Pajtler, K.
Kalish, J. M. ; Becktell, K. D. ; Bougeard, G. ; Brodeur, G. M. ; Diller, L. R. ; Doria, A. S. ; Hansford, J. R. ; Klein, S. D. ; Kohlmann, W. K. ; Kratz, C. P. ; MacFarland, S. P. ; Pajtler, K. ; Rednam, S. P. ; Schienda, J. ; States, L. J. ; Villani, A. ; Weksberg, R. ; Zelley, K. ; Tomlinson, G. E. ; Brzezinski, J. J.
Genard, G. ; Tirinato, L. ; Pagliari, F. ; Da Silva, J. ; Giammona, A. ; Alquraish, F. ; Reyes, M. P. ; Bordas, M. ; Marafioti, M. G. ; Franco, S. D. ; Janssen, J. ; Garcia-Calderón, D. ; Hanley, R. ; Nistico, C. ; Fukasawa, Y. ; Müller, T. ; Krijgsveld, J. ; Todaro, M. ; Costanzo, F. S. ; Stassi, G. ; Nessling, M. ; Richter, K. ; Maass, K. K. ; Liberale, C. ; Seco, J.
Koedijk, J. B. ; van der Werf, I. ; Penter, L. ; Vermeulen, M. A. ; Barneh, F. ; Perzolli, A. ; Meesters-Ensing, J. I. ; Metselaar, D. S. ; Margaritis, T. ; Fiocco, M. ; de Groot-Kruseman, H. A. ; Moeniralam, R. ; Bang Christensen, K. ; Porter, B. ; Pfaff, K. ; Garcia, J. S. ; Rodig, S. J. ; Wu, C. J. ; Hasle, H. ; Nierkens, S. ; Belderbos, M. E. ; Zwaan, C. M. ; Heidenreich, O.
Hansford, J. R. ; Bouche, G. ; Ramaswamy, V. ; Jabado, N. ; Fonseca, A. ; Moloney, S. ; Gottardo, N. G. ; Robinson, G. W. ; Gajjar, A. ; Tinkle, C. L. ; Fisher, P. G. ; Foreman, N. ; Ashley, D. M. ; Ziegler, D. S. ; Eisenstat, D. D. ; Massimino, M. ; Witt, O. ; Bartels, U. ; Rutkowski, S. ; Hargrave, D. ; Fouladi, M. ; Pfister, S. M. ; Bouffet, E.
Went, M. ; Duran-Lozano, L. ; Halldorsson, G. H. ; Gunnell, A. ; Ugidos-Damboriena, N. ; Law, P. ; Ekdahl, L. ; Sud, A. ; Thorleifsson, G. ; Thodberg, M. ; Olafsdottir, T. ; Lamarca-Arrizabalaga, A. ; Cafaro, C. ; Niroula, A. ; Ajore, R. ; Lopez de Lapuente Portilla, A. ; Ali, Z. ; Pertesi, M. ; Goldschmidt, H. ; Stefansdottir, L. ; Kristinsson, S. Y. ; Stacey, S. N. ; Love, T. J. ; Rognvaldsson, S. ; Hajek, R. ; Vodicka, P. ; Pettersson-Kymmer, U. ; Späth, F. ; Schinke, C. ; Van Rhee, F. ; Sulem, P. ; Ferkingstad, E. ; Hjorleifsson Eldjarn, G. ; Mellqvist, U.-H. ; Jonsdottir, I. ; Morgan, G. ; Sonneveld, P. ; Waage, A. ; Weinhold, N. ; Thomsen, H. ; Försti, A. ; Hansson, M. ; Juul-Vangsted, A. ; Thorsteinsdottir, U. ; Hemminki, K. ; Kaiser, M. ; Rafnar, T. ; Stefansson, K. ; Houlston, R. ; Nilsson, B.
Godbole, S. ; Voß, H. ; Gocke, A. ; Schlumbohm, S. ; Schumann, Y. ; Peng, B. ; Mynarek, M. ; Rutkowski, S. ; Dottermusch, M. ; Dorostkar, M. M. ; Korshunov, A. ; Mair, T. ; Pfister, S. ; Kwiatkowski, M. ; Hotze, M. ; Neumann, P. ; Hartmann, C. ; Weis, J. ; Liesche-Starnecker, F. ; Guan, Y. ; Moritz, M. ; Siebels, B. ; Struve, N. ; Schlüter, H. ; Schüller, U. ; Krisp, C. ; Neumann, J. E.
Sievers, P. ; Bielle, F. ; Göbel, K. ; Schrimpf, D. ; Nichelli, L. ; Mathon, B. ; Appay, R. ; Boldt, H. B. ; Dohmen, H. ; Selignow, C. ; Acker, T. ; Vicha, A. ; Martinetto, H. ; Schweizer, L. ; Schüller, U. ; Brandner, S. ; Wesseling, P. ; Schmid, S. ; Capper, D. ; Abdullaev, Z. ; Aldape, K. ; Korshunov, A. ; Krieg, S. M. ; Wick, W. ; Pfister, S. M. ; von Deimling, A. ; Reuss, D. E. ; Jones, D. ; Sahm, F.
Adolph, J. E. ; Fleischhack, G. ; Tschirner, S. ; Rink, L. ; Dittes, C. ; Mikasch, R. ; Dammann, P. ; Mynarek, M. ; Obrecht-Sturm, D. ; Rutkowski, S. ; Bison, B. ; Warmuth-Metz, M. ; Pietsch, T. ; Pfister, S. ; Pajtler, K. ; Milde, T. ; Kortmann, R.-D. ; Dietzsch, S. ; Timmermann, B. ; Tippelt, S. ; HIT-Network, G. G.
Zuckermann, M. ; He, C. ; Andrews, J. ; Bagchi, A. ; Sloan-Henry, R. ; Bianski, B. ; Xie, J. ; Wang, Y. ; Twarog, N. ; Onar-Thomas, A. ; Ernst, K. ; Yang, L. ; Li, Y. ; Zhu, X. ; Ocasio, J. K. ; Budd, K. M. ; Dalton, J. ; Li, X. ; Chepyala, D. ; Zhang, J. ; Xu, K. ; Hover, L. ; Roach, J. T. ; Chan, K. C. ; Hofmann, N. ; McKinnon, P. J. ; Pfister, S. ; Shelat, A. A. ; Rankovic, Z. ; Freeman, B. B. ; Chiang, J. ; Jones, D. ; Tinkle, C. L. ; Baker, S. J.
Shiraishi, R. ; Cancila, G. ; Kumegawa, K. ; Torrejon, J. ; Basili, I. ; Bernardi, F. ; Silva, P. B. G. d. ; Wang, W. ; Chapman, O. ; Yang, L. ; Jami, M. ; Nishitani, K. ; Arai, Y. ; Xiao, Z. ; Yu, H. ; Lo Re, V. ; Marsaud, V. ; Talbot, J. ; Lombard, B. ; Loew, D. ; Jingu, M. ; Okonechnikov, K. ; Sone, M. ; Motohashi, N. ; Aoki, Y. ; Pfister, S. M. ; Chavez, L. ; Hoshino, M. ; Maruyama, R. ; Ayrault, O. ; Kawauchi, D.
Mynarek, M. ; Rossius, A. ; Guiard, A. ; Ottensmeier, H. ; von Hoff, K. ; Obrecht-Sturm, D. ; Bußenius, L. ; Friedrich, C. ; von Bueren, A. O. ; Gerber, N. U. ; Traunwieser, T. ; Kortmann, R.-D. ; Warmuth-Metz, M. ; Bison, B. ; Thomale, U.-W. ; Krauss, J. ; Pietsch, T. ; Clifford, S. C. ; Pfister, S. M. ; Sturm, D. ; Sahm, F. ; Tischler, T. ; Rutkowski, S.
Kocher, D. ; Cao, L. ; Guiho, R. ; Langhammer, M. ; Lai, Y.-L. ; Becker, P. ; Hamdi, H. ; Friedel, D. ; Selt, F. ; Vonhören, D. ; Zaman, J. ; Valinciute, G. ; Herter, S. ; Picard, D. ; Rettenmeier, J. ; Maass, K. K. ; Pajtler, K. W. ; Remke, M. ; von Deimling, A. ; Pusch, S. ; Pfister, S. M. ; Oehme, I. ; Jones, D. T. W. ; Halbach, S. ; Brummer, T. ; Martinez-Barbera, J. P. ; Witt, O. ; Milde, T. ; Sigaud, R.
Hench, J. ; Hultschig, C. ; Brugger, J. ; Mariani, L. ; Guzman, R. ; Soleman, J. ; Leu, S. ; Benton, M. ; Stec, I. M. ; Hench, I. B. ; Hoffmann, P. ; Harter, P. ; Weber, K. ; Albers, A. ; Thomas, C. ; Hasselblatt, M. ; Schüller, U. ; Restelli, L. ; Capper, D. ; Hewer, E. ; Diebold, J. ; Kolenc, D. ; Schneider, U. C. ; Rushing, E. ; Della Monica, R. ; Chiariotti, L. ; Sill, M. ; Schrimpf, D. ; von Deimling, A. ; Sahm, F. ; Kölsche, C. ; Tolnay, M. ; Frank, S.
Gilbertson, R. J. ; Behjati, S. ; Böttcher, A.-L. ; Bronner, M. E. ; Burridge, M. ; Clausing, H. ; Clifford, H. ; Danaher, T. ; Donovan, L. K. ; Drost, J. ; Eggermont, A. M. M. ; Emerson, C. ; Flores, M. G. ; Hamerlik, P. ; Jabado, N. ; Jones, A. ; Kaessmann, H. ; Kleinman, C. L. ; Kool, M. ; Kutscher, L. ; Lindberg, G. ; Linnane, E. ; Marioni, J. C. ; Maris, J. M. ; Monje, M. ; Macaskill, A. ; Niederer, S. ; Northcott, P. A. ; Peeters, E. ; Plieger-van Solkema, W. ; Preußner, L. ; Rios, A. C. ; Rippe, K. ; Sandford, P. ; Sgourakis, N. G. ; Shlien, A. ; Smith, P. ; Straathof, K. ; Sullivan, P. J. ; Suvà, M. L. ; Taylor, M. D. ; Thompson, E. ; Vento-Tormo, R. ; Wainwright, B. J. ; Wechsler-Reya, R. J. ; Westermann, F. ; Winslade, S. ; Al-Lazikani, B. ; Pfister, S.
Trkova, K. ; Sumerauer, D. ; Bubenikova, A. ; Krskova, L. ; Vicha, A. ; Koblizek, M. ; Zamecnik, J. ; Jurasek, B. ; Kyncl, M. ; Malinova, B. ; Ondrova, B. ; Jones, D. T. W. ; Sill, M. ; Strnadova, M. ; Stolova, L. ; Misove, A. ; Benes, V. ; Zapotocky, M.
Iser, F. ; Hinz, F. ; Hoffmann, D. C. F. ; Grassl, N. ; Güngör, C. ; Meyer, J. ; Dörner, L. ; Hofmann, L. ; Kelbch, V. ; Göbel, K. ; Mahmutoglu, M. A. ; Vollmuth, P. ; Patel, A. J. ; Nguyen, D. ; Kaulen, L. ; Mildenberger, I. ; Sahm, K. ; Maass, K. ; Pajtler, K. ; Shankar, G. M. ; Weiler, M. ; Wildemann, B. ; Winkler, F. ; von Deimling, A. ; Platten, M. ; Wick, W. ; Sahm, F. ; Kessler, T.
Sprokkerieft, J. ; van der Beek, J. N. ; Spreafico, F. ; Selle, B. ; Thebaud, E. ; Chowdhury, T. ; Brok, J. ; Ottóffy, G. ; Sun, X. ; Ramírez Villar, G. L. ; Sagoyan, G. ; Segers, H. ; Doganis, D. ; Serra, A. ; Lemelle, L. ; Graf, N. ; Verschuur, A. C. ; Tytgat, G. A. M. ; van den Heuvel-Eibrink, M. M.
Gödicke, S. ; Kresbach, C. ; Ehlert, M. ; Obrecht, D. ; Altendorf, L. ; Hack, K. ; von Hoff, K. ; Carén, H. ; Melcher, V. ; Kerl, K. ; Englinger, B. ; Filbin, M. ; Pajtler, K. W. ; Gojo, J. ; Pietsch, T. ; Rutkowski, S. ; Schüller, U.
Daenekas, B. ; Pérez, E. ; Boniolo, F. ; Stefan, S. ; Benfatto, S. ; Sill, M. ; Sturm, D. ; Jones, D. T. W. ; Capper, D. ; Zapatka, M. ; Hovestadt, V.
Ghasemi, D. R. ; Okonechnikov, K. ; Rademacher, A. ; Tirier, S. ; Maass, K. K. ; Schumacher, H. ; Joshi, P. ; Gold, M. P. ; Sundheimer, J. ; Statz, B. ; Rifaioglu, A. S. ; Bauer, K. ; Schumacher, S. ; Bortolomeazzi, M. ; Giangaspero, F. ; Ernst, K. ; Clifford, S. C. ; Saez-Rodriguez, J. ; Jones, D. ; Kawauchi, D. ; Fraenkel, E. ; Mallm, J.-P. ; Rippe, K. ; Korshunov, A. ; Pfister, S. ; Pajtler, K.
Gold, M. P. ; Ong, W. ; Masteller, A. M. ; Ghasemi, D. R. ; Galindo, J. A. ; Park, N. R. ; Huynh, N. C. ; Donde, A. ; Pister, V. ; Saurez, R. A. ; Vladoiu, M. C. ; Hwang, G. H. ; Eisemann, T. ; Donovan, L. K. ; Walker, A. D. ; Benetatos, J. ; Dufour, C. ; Garzia, L. ; Segal, R. A. ; Wechsler-Reya, R. J. ; Mesirov, J. P. ; Korshunov, A. ; Pajtler, K. ; Pomeroy, S. L. ; Ayrault, O. ; Davidson, S. M. ; Cotter, J. A. ; Taylor, M. D. ; Fraenkel, E.
Sepp, M. ; Leiss, K. ; Murat, F. ; Okonechnikov, K. ; Joshi, P. K. ; Leushkin, E. ; Spänig, L. ; Mbengue, N. ; Schneider, C. ; Schmidt, J. ; Trost, N. ; Schauer, M. ; Khaitovich, P. ; Lisgo, S. ; Palkovits, M. ; Giere, P. ; Kutscher, L. ; Anders, S. ; Cardoso-Moreira, M. ; Sarropoulos, I. ; Pfister, S. ; Kaessmann, H.
Tonn, S. ; Korshunov, A. ; Obrecht, D. ; Sill, M. ; Spohn, M. ; von Hoff, K. ; Milde, T. ; Pietsch, T. ; Goschzik, T. ; Bison, B. ; Juhnke, B.-O. ; Struve, N. ; Sturm, D. ; Sahm, F. ; Bockmayr, M. ; Friedrich, C. ; von Bueren, A. O. ; Gerber, N. U. ; Benesch, M. ; Jones, D. ; Kool, M. ; Wefers, A. K. ; Schüller, U. ; Pfister, S. ; Rutkowski, S. ; Mynarek, M.
Arseni, L. ; Sharma, R. ; Mack, N. ; Nagalla, D. ; Ohl, S. ; Hielscher, T. ; Singhal, M. ; Engel, R. ; Augustin, H. ; Sandhoff, R. ; Herold-Mende, C. ; Tews, B. ; Lichter, P. ; Seiffert, M.
Niazi, Y. ; Paramasivam, N. ; Blocka, J. ; Kumar, A. ; Huhn, S. ; Schlesner, M. ; Weinhold, N. ; Sijmons, R. ; De Jong, M. ; Durie, B. ; Goldschmidt, H. ; Hemminki, K. ; Försti, A.
Ecker, J. ; Selt, F. ; Sturm, D. ; Sill, M. ; Korshunov, A. ; Hirsch, S. ; Capper, D. ; Dikow, N. ; Sutter, C. ; Müller, C. ; Sigaud, R. ; Eggert, A. ; Simon, T. ; Niehues, T. ; von Deimling, A. ; Pajtler, K. W. ; van Tilburg, C. M. ; Jones, D. T. W. ; Sahm, F. ; Pfister, S. M. ; Witt, O. ; Milde, T.
Mynarek, M. ; Obrecht, D. ; Sill, M. ; Sturm, D. ; Kloth-Stachnau, K. ; Selt, F. ; Ecker, J. ; von Hoff, K. ; Juhnke, B.-O. ; Goschzik, T. ; Pietsch, T. ; Bockmayr, M. ; Kool, M. ; von Deimling, A. ; Witt, O. ; Schüller, U. ; Benesch, M. ; Gerber, N. U. ; Sahm, F. ; Jones, D. ; Korshunov, A. ; Pfister, S. ; Rutkowski, S. ; Milde, T.
Selt, F. ; Sigaud, R. ; Valinciute, G. ; Sievers, P. ; Zaman, J. ; Alco, C. ; Schmid, S. ; Peterziel, H. ; Tsai, J. W. ; Guiho, R. ; Martínez-Barbera, J. P. ; Pusch, S. ; Deng, J. ; Zhai, Y. ; van Tilburg, C. M. ; Schuhman, M. U. ; Damaty, A. E. L. ; Bandopadhayay, P. ; Herold-Mende, C. ; von Deimling, A. ; Pfister, S. ; Montero, J. ; Capper, D. ; Oehme, I. ; Sahm, F. ; Jones, D. ; Witt, O. ; Milde, T.
Sigaud, R. ; Rösch, L. ; Gatzweiler, C. ; Benzel, J. ; von Soosten, L. ; Peterziel, H. ; Selt, F. ; Najafi, S. ; Ayhan, S. ; Gerloff, X. ; Hofmann, N. ; Büdenbender, I. ; Schmitt, L. ; Foerster, K. I. ; Burhenne, J. ; Haefeli, W. E. ; Korshunov, A. ; Sahm, F. ; van Tilburg, C. M. ; Jones, D. ; Pfister, S. ; Knoerzer, D. ; Kreider, B. L. ; Sauter, M. ; Pajtler, K. ; Zuckermann, M. ; Oehme, I. ; Witt, O. ; Milde, T.
Lago, C. ; Federico, A. ; Leva, G. ; Mack, N. L. ; Schwalm, B. ; Ballabio, C. ; Gianesello, M. ; Abballe, L. ; Giovannoni, I. ; Reddel, S. ; Rossi, S. ; Leone, N. ; Carai, A. ; Mastronuzzi, A. ; Bisio, A. ; Soldano, A. ; Quintarelli, C. ; Locatelli, F. ; Kool, M. ; Miele, E. ; Tiberi, L.
Selt, F. ; El Damaty, A. ; Schuhmann, M. U. ; Sigaud, R. ; Ecker, J. ; Sievers, P. ; Kocher, D. ; Herold-Mende, C. ; Oehme, I. ; von Deimling, A. ; Pfister, S. ; Sahm, F. ; Jones, D. ; Witt, O. ; Milde, T.
Schoof, M. ; Godbole, S. ; Albert, T. K. ; Dottermusch, M. ; Walter, C. ; Ballast, A. ; Qin, N. ; Olivera, M. B. ; Göbel, C. ; Neyazi, S. ; Holdhof, D. ; Kresbach, C. ; Peter, L.-S. ; Epplen, G. D. ; Thaden, V. ; Spohn, M. ; Blattner-Johnson, M. ; Modemann, F. ; Mynarek, M. ; Rutkowski, S. ; Sill, M. ; Varghese, J. ; Afflerbach, A.-K. ; Eckhardt, A. ; Münter, D. ; Verma, A. ; Struve, N. ; Jones, D. ; Remke, M. ; Neumann, J. E. ; Kerl, K. ; Schüller, U.
Rautajoki, K. J. ; Jaatinen, S. ; Hartewig, A. ; Tiihonen, A. M. ; Annala, M. ; Salonen, I. ; Valkonen, M. ; Simola, V. ; Vuorinen, E. M. ; Kivinen, A. ; Rauhala, M. J. ; Nurminen, R. ; Maass, K. K. ; Lahtela, S.-L. ; Jukkola, A. ; Yli-Harja, O. ; Helén, P. ; Pajtler, K. W. ; Ruusuvuori, P. ; Haapasalo, J. ; Zhang, W. ; Haapasalo, H. ; Nykter, M.
Peyrl, A. ; Chocholous, M. ; Sabel, M. ; Lassaletta, A. ; Sterba, J. ; Leblond, P. ; Nysom, K. ; Torsvik, I. ; Chi, S. N. ; Perwein, T. ; Jones, N. ; Holm, S. ; Nyman, P. ; Mörse, H. ; Öberg, A. ; Weiler-Wichtl, L. ; Leiss, U. ; Haberler, C. ; Schmook, M. T. ; Mayr, L. ; Dieckmann, K. ; Kool, M. ; Gojo, J. ; Azizi, A. A. ; André, N. ; Kieran, M. ; Slavc, I.
Ciobanu-Caraus, O. ; Czech, T. ; Peyrl, A. ; Haberler, C. ; Kasprian, G. ; Furtner, J. ; Kool, M. ; Sill, M. ; Frischer, J. M. ; Cho, A. ; Slavc, I. ; Rössler, K. ; Gojo, J. ; Dorfer, C.
Nabbi, A. ; Beck, P. ; Delaidelli, A. ; Oldridge, D. A. ; Sudhaman, S. ; Zhu, K. ; Yang, S. Y. C. ; Mulder, D. T. ; Bruce, J. P. ; Paulson, J. N. ; Raman, P. ; Zhu, Y. ; Resnick, A. C. ; Sorensen, P. H. ; Sill, M. ; Brabetz, S. ; Lambo, S. ; Malkin, D. ; Johann, P. ; Kool, M. ; Jones, D. ; Pfister, S. ; Jäger, N. ; Pugh, T. J.
Okonechnikov, K. ; Joshi, P. K. ; Sepp, M. ; Leiss, K. ; Sarropoulos, I. ; Murat, F. ; Sill, M. ; Beck, P. ; Chan, K. C. ; Korshunov, A. ; Sahm, F. ; Deng, M. Y. ; Sturm, D. ; DeSisto, J. ; Donson, A. M. ; Foreman, N. K. ; Green, A. L. ; Robinson, G. ; Orr, B. A. ; Gao, Q. ; Darrow, E. ; Hadley, J. L. ; Northcott, P. A. ; Gojo, J. ; Kawauchi, D. ; Hovestadt, V. ; Filbin, M. G. ; von Deimling, A. ; Zuckermann, M. ; Pajtler, K. ; Kool, M. ; Jones, D. ; Jäger, N. ; Kutscher, L. ; Kaessmann, H. ; Pfister, S.
White, C. L. ; Kinross, K. M. ; Buntine, M. K. ; Rasouli, E. ; Strong, R. ; Jones, J. M. ; Cain, J. E. ; Sturm, D. ; Sahm, F. ; Jones, D. T. W. ; Pfister, S. M. ; Robertson, T. ; D'Arcy, C. ; Rodriguez, M. L. ; Dyke, J. M. ; Junckerstorff, R. ; Bhuva, D. D. ; Davis, M. J. ; Wood, P. ; Hassall, T. ; Ziegler, D. S. ; Kellie, S. ; McCowage, G. ; Alvaro, F. ; Kirby, M. ; Heath, J. A. ; Tsui, K. ; Dodgshun, A. ; Eisenstat, D. D. ; Khuong-Quang, D.-A. ; Wall, M. ; Algar, E. M. ; Gottardo, N. G. ; Hansford, J. R.
Sigaud, R. ; Albert, T. K. ; Hess, C. ; Hielscher, T. ; Winkler, N. ; Kocher, D. ; Walter, C. ; Münter, D. ; Selt, F. ; Usta, D. ; Ecker, J. ; Brentrup, A. ; Hasselblatt, M. ; Thomas, C. ; Varghese, J. ; Capper, D. ; Thomale, U. W. ; Hernáiz Driever, P. ; Simon, M. ; Horn, S. ; Herz, N. A. ; Koch, A. ; Sahm, F. ; Hamelmann, S. ; Faria-Andrade, A. ; Jabado, N. ; Schuhmann, M. U. ; Schouten-van Meeteren, A. Y. N. ; Hoving, E. ; Brummer, T. ; van Tilburg, C. M. ; Pfister, S. M. ; Witt, O. ; Jones, D. ; Kerl, K. ; Milde, T.
Funke, V. L. E. ; Sandmann, S. ; Melcher, V. ; Seggewiss, J. ; Horvath, J. ; Jäger, N. ; Kool, M. ; Jones, D. T. W. ; Pfister, S. M. ; Milde, T. ; Rutkowski, S. ; Mynarek, M. ; Varghese, J. ; Sträter, R. ; Rust, S. ; Seelhöfer, A. ; Reunert, J. ; Fiedler, B. ; Schüller, U. ; Marquardt, T. ; Kerl, K.
Johann, P. ; Altendorf, L. ; Efremova, E.-M. ; Holsten, T. ; Steinbügl, M. ; Nemes, K. ; Eckhardt, A. ; Kresbach, C. ; Bockmayr, M. ; Koch, A. ; Haberler, C. ; Antonelli, M. ; DeSisto, J. ; Schuhmann, M. U. ; Hauser, P. ; Siebert, R. ; Bens, S. ; Kool, M. ; Green, A. L. ; Hasselblatt, M. ; Frühwald, M. C. ; Schüller, U.
Heipertz, A.-E. ; Pajtler, K. ; Pfaff, E. ; Schramm, K. ; Blattner-Johnson, M. ; Milde, T. ; Jones, B. ; Zuliani, C. ; Hutter, C. ; Lohi, O. ; Kattamis, A. ; Dachowska-Kalwak, I. ; Nilsson, A. ; Gerber, N. U. ; Langenberg, K. P. S. ; Goemans, B. ; Zwaan, C. M. ; Molenaar, J. J. ; Jäger, N. ; Dirksen, U. ; Witt, R. ; Pfister, S. ; Jones, D. ; Kopp-Schneider, A. ; Witt, O. ; van Tilburg, C. M.
Ng, C. H. ; Obrecht, D. ; Wells, O. ; Zapotocky, M. ; Sumerauer, D. ; Coltin, H. ; Khuong-Quang, D.-A. ; Eisenstat, D. D. ; Kinross, K. M. ; White, C. L. ; Algar, E. M. ; Luck, A. ; Witt, H. ; Schüller, U. ; Mynarek, M. ; Pietsch, T. ; Gerber, N. U. ; Benesch, M. ; Warmuth-Metz, M. ; Kortmann, R. ; Bison, B. ; Taylor, M. D. ; Rutkowski, S. ; Pfister, S. M. ; Jones, D. T. ; Gottardo, N. G. ; von Hoff, K. ; Pajtler, K. ; Ramaswamy, V. ; Hansford, J. R.
Deng, M. Y. ; Hinz, F. ; Maas, S. L. N. ; Anil, G. ; Sievers, P. ; Conde-Lopez, C. ; Lischalk, J. ; Rauh, S. ; Eichkorn, T. ; Regnery, S. ; Bauer, L. ; Held, T. ; Meixner, E. ; Lang, K. ; Hörner-Rieber, J. ; Herfarth, K. ; Jones, D. ; Pfister, S. ; Jungk, C. ; Unterberg, A. ; Wick, W. ; von Deimling, A. ; Debus, J. ; Sahm, F. ; König, L.
Funke, V. L. E. ; Walter, C. ; Melcher, V. ; Wei, L. ; Sandmann, S. ; Hotfilder, M. ; Varghese, J. ; Jäger, N. ; Kool, M. ; Jones, D. T. W. ; Pfister, S. M. ; Milde, T. ; Mynarek, M. ; Rutkowski, S. ; Seggewiss, J. ; Jeising, D. ; de Faria, F. W. ; Marquardt, T. ; Albert, T. K. ; Schüller, U. ; Kerl, K.
Donson, A. M. ; Bertrand, K. C. ; Riemondy, K. A. ; Gao, D. ; Zhuang, Y. ; Sanford, B. ; Norris, G. A. ; Chapman, R. J. ; Fu, R. ; Willard, N. ; Griesinger, A. M. ; de Sousa, G. R. ; Amani, V. ; Grimaldo, E. ; Hankinson, T. C. ; Booker, F. ; Sill, M. ; Grundy, R. G. ; Pajtler, K. ; Ellison, D. W. ; Foreman, N. K. ; Ritzmann, T. A.
Deng, M. ; Hinz, F. ; Harrabi, S. ; Sturm, D. ; Sill, M. ; Korshunov, A. ; Eichkorn, T. ; Hörner-Rieber, J. ; Herfarth, K. ; Jungk, C. ; Unterberg, A. ; Pfister, S. ; Wick, W. ; von Deimling, A. ; Jones, D. ; Debus, J. ; Sahm, F. ; König, L.
Valinciute, G. ; Ecker, J. ; Selt, F. ; Hielscher, T. ; Sigaud, R. ; Ridinger, J. ; Thatikonda, V. ; Gatzweiler, C. ; Robinson, S. ; Talbot, J. ; Bernardi, F. ; Picard, D. ; Blattner-Johnson, M. ; Schmid, S. ; Jones, D. T. ; van Tilburg, C. M. ; Capper, D. ; Kool, M. ; Remke, M. ; Oehme, I. ; Pfister, S. M. ; Roussel, M. F. ; Ayrault, O. ; Witt, O. ; Milde, T.
Cabrera-Serrano, A. J. ; Sánchez-Maldonado, J. M. ; Ter Horst, R. ; Macauda, A. ; García-Martín, P. ; Benavente, Y. ; Landi, S. ; Clay-Gilmour, A. ; Niazi, Y. ; Espinet, B. ; Rodríguez-Sevilla, J. J. ; Pérez, E. M. ; Maffei, R. ; Blanco, G. ; Giaccherini, M. ; Cerhan, J. R. ; Marasca, R. ; López-Nevot, M. Á. ; Chen-Liang, T. ; Thomsen, H. ; Gámez, I. ; Campa, D. ; Moreno, V. ; de Sanjosé, S. ; Marcos-Gragera, R. ; García-Álvarez, M. ; Dierssen-Sotos, T. ; Jerez, A. ; Butrym, A. ; Norman, A. D. ; Luppi, M. ; Slager, S. L. ; Hemminki, K. ; Li, Y. ; Berndt, S. I. ; Casabonne, D. ; Alcoceba, M. ; Puiggros, A. ; Netea, M. G. ; Försti, A. ; Canzian, F. ; Sainz, J.
Korshunov, A. ; Okonechnikov, K. ; Schrimpf, D. ; Tonn, S. ; Mynarek, M. ; Koster, J. ; Sievers, P. ; Milde, T. ; Sahm, F. ; Jones, D. ; von Deimling, A. ; Pfister, S. ; Kool, M.
Hardin, E. C. ; Schmid, S. ; Sommerkamp, A. ; Bodden, C. ; Heipertz, A.-E. ; Sievers, P. ; Wittmann, A. ; Milde, T. ; Pfister, S. ; von Deimling, A. ; Horn, S. ; Herz, N. A. ; Simon, M. ; Perera, A. A. ; Azizi, A. ; Cruz, O. ; Curry, S. ; Van Damme, A. ; Garami, M. ; Hargrave, D. ; Kattamis, A. ; Kotnik, B. F. ; Lähteenmäki, P. ; Scheinemann, K. ; Schouten-van Meeteren, A. Y. N. ; Sehested, A. ; Viscardi, E. ; Wormdal, O. M. ; Zapotocky, M. ; Ziegler, D. S. ; Koch, A. ; Driever, P. H. ; Witt, O. ; Capper, D. ; Sahm, F. ; Jones, D. ; van Tilburg, C. M.
Liu, Y. ; Klein, J. ; Bajpai, R. ; Dong, L. ; Tran, Q. ; Kolekar, P. ; Smith, J. L. ; Ries, R. E. ; Huang, B. J. ; Wang, Y.-C. ; Alonzo, T. A. ; Tian, L. ; Mulder, H. L. ; Shaw, T. I. ; Ma, J. ; Walsh, M. P. ; Song, G. ; Westover, T. ; Autry, R. J. ; Gout, A. M. ; Wheeler, D. A. ; Wan, S. ; Wu, G. ; Yang, J. J. ; Evans, W. E. ; Loh, M. ; Easton, J. ; Zhang, J. ; Klco, J. M. ; Meshinchi, S. ; Brown, P. A. ; Pruett-Miller, S. M. ; Ma, X.
Moreno, L. ; DuBois, S. G. ; Glade Bender, J. ; Mauguen, A. ; Bird, N. ; Buenger, V. ; Casanova, M. ; Doz, F. ; Fox, E. ; Gore, L. ; Hawkins, D. S. ; Izraeli, S. ; Jones, D. ; Kearns, P. R. ; Molenaar, J. J. ; Nysom, K. ; Pfister, S. ; Reaman, G. ; Smith, M. ; Weigel, B. ; Vassal, G. ; Zwaan, C. M. ; Paoletti, X. ; Iasonos, A. ; Pearson, A. D. J.
Goddard, J. ; Castle, J. ; Southworth, E. ; Fletcher, A. ; Crosier, S. ; Martin-Guerrero, I. ; García-Ariza, M. ; Navajas, A. ; Masliah-Planchon, J. ; Bourdeaut, F. ; Dufour, C. ; Ayrault, O. ; Goschzik, T. ; Pietsch, T. ; Sill, M. ; Pfister, S. ; Rutkowski, S. ; Richardson, S. ; Hill, R. M. ; Williamson, D. ; Bailey, S. ; Schwalbe, E. C. ; Clifford, S. C. ; Hicks, D.
Sievers, P. ; Sill, M. ; Schrimpf, D. ; Abdullaev, Z. ; Donson, A. M. ; Lake, J. A. ; Friedel, D. ; Scheie, D. ; Tynninen, O. ; Rauramaa, T. ; Vepsäläinen, K. L. ; Samuel, D. ; Chapman, R. ; Grundy, R. G. ; Pajtler, K. W. ; Tauziède-Espariat, A. ; Métais, A. ; Varlet, P. ; Snuderl, M. ; Jacques, T. S. ; Aldape, K. ; Reuss, D. E. ; Korshunov, A. ; Wick, W. ; Pfister, S. M. ; von Deimling, A. ; Sahm, F. ; Jones, D.
Ecker, J. ; Selt, F. ; Sturm, D. ; Sill, M. ; Korshunov, A. ; Hirsch, S. ; Capper, D. ; Dikow, N. ; Sutter, C. ; Müller, C. ; Sigaud, R. ; Eggert, A. ; Simon, T. ; Niehues, T. ; von Deimling, A. ; Pajtler, K. ; van Tilburg, C. M. ; Jones, D. ; Sahm, F. ; Pfister, S. ; Witt, O. ; Milde, T.
Chapman, R. J. ; Ghasemi, D. R. ; Andreiuolo, F. ; Zschernack, V. ; Tauziede Espariat, A. ; Buttarelli, F. R. ; Giangaspero, F. ; Grill, J. ; Haberler, C. ; Paine, S. M. L. ; Scott, I. ; Jacques, T. S. ; Sill, M. ; Pfister, S. ; Kilday, J.-P. ; Leblond, P. ; Massimino, M. ; Witt, H. ; Modena, P. ; Varlet, P. ; Pietsch, T. ; Grundy, R. G. ; Pajtler, K. ; Ritzmann, T. A. ; Childhood, B. o. E. i. ; Adolescence
Hemminki, K. ; Li, X. ; Försti, A. ; Eng, C.
Tauziède-Espariat, A. ; Uro-Coste, E. ; Sievers, P. ; Nicaise, Y. ; Mariet, C. ; Siegfried, A. ; Pierron, G. ; Guillemot, D. ; Benzakoun, J. ; Pallud, J. ; Roques, M. ; Bonneville, F. ; Larrieu-Ciron, D. ; Chaynes, P. ; Saffroy, R. ; Hamelin, J. ; Hasty, L. ; Métais, A. ; Chrétien, F. ; Kool, M. ; Gojo, J. ; Varlet, P. ; RENOCLIP-LOC
Pun, M. ; Pratt, D. ; Nano, P. R. ; Joshi, P. K. ; Jiang, L. ; Englinger, B. ; Rao, A. ; Cieslik, M. ; Chinnaiyan, A. M. ; Aldape, K. ; Pfister, S. ; Filbin, M. G. ; Bhaduri, A. ; Venneti, S.
Luo, Z. ; Xin, D. ; Liao, Y. ; Berry, K. ; Ogurek, S. ; Zhang, F. ; Zhang, L. ; Zhao, C. ; Rao, R. ; Dong, X. ; Li, H. ; Yu, J. ; Lin, Y. ; Huang, G. ; Xu, L. ; Xin, M. ; Nishinakamura, R. ; Yu, J. ; Kool, M. ; Pfister, S. M. ; Roussel, M. F. ; Zhou, W. ; Weiss, W. A. ; Andreassen, P. ; Lu, Q. R.
Peterziel, H. ; Jamaladdin, N. ; ElHarouni, D. ; Gerloff, X. ; Herter, S. ; Fiesel, P. ; Berker, Y. ; Blattner-Johnson, M. ; Schramm, K. ; Jones, B. ; Reuss, D. ; Turunen, L. ; Friedenauer, A. ; Holland-Letz, T. ; Sill, M. ; Weiser, L. ; Previti, C. ; Balasubramanian, G. ; Gerber, N. U. ; Gojo, J. ; Hutter, C. ; Øra, I. ; Lohi, O. ; Kattamis, A. ; de Wilde, B. ; Westermann, F. ; Tippelt, S. ; Graf, N. ; Nathrath, M. ; Sparber-Sauer, M. ; Sehested, A. ; Kramm, C. M. ; Dirksen, U. ; Kallioniemi, O. ; Pfister, S. ; van Tilburg, C. M. ; Jones, D. ; Saarela, J. ; Pietiäinen, V. ; Jäger, N. ; Schlesner, M. ; Kopp-Schneider, A. ; Oppermann, S. ; Milde, T. ; Witt, O. ; Oehme, I.
Ajore, R. ; Niroula, A. ; Pertesi, M. ; Cafaro, C. ; Thodberg, M. ; Went, M. ; Bao, E. L. ; Duran-Lozano, L. ; Lopez de Lapuente Portilla, A. ; Olafsdottir, T. ; Ugidos-Damboriena, N. ; Magnusson, O. ; Samur, M. ; Lareau, C. A. ; Halldorsson, G. H. ; Thorleifsson, G. ; Norddahl, G. L. ; Gunnarsdottir, K. ; Försti, A. ; Goldschmidt, H. ; Hemminki, K. ; van Rhee, F. ; Kimber, S. ; Sperling, A. S. ; Kaiser, M. ; Anderson, K. ; Jonsdottir, I. ; Munshi, N. ; Rafnar, T. ; Waage, A. ; Weinhold, N. ; Thorsteinsdottir, U. ; Sankaran, V. G. ; Stefansson, K. ; Houlston, R. ; Nilsson, B.
Orth, M. F. ; Surdez, D. ; Faehling, T. ; Ehlers, A. ; Marchetto, A. ; Grossetête, S. ; Volckmann, R. ; Zwijnenburg, D. A. ; Gerke, J. S. ; Zaidi, S. ; Alonso, J. ; Sastre, A. ; Baulande, S. ; Sill, M. ; Cidre-Aranaz, F. ; Ohmura, S. ; Kirchner, T. ; Hauck, S. M. ; Reischl, E. ; Gymrek, M. ; Pfister, S. ; Strauch, K. ; Koster, J. ; Delattre, O. ; Grünewald, T.
Brigante, G. ; Lazzaretti, C. ; Paradiso, E. ; Nuzzo, F. ; Sitti, M. ; Tüttelmann, F. ; Moretti, G. ; Silvestri, R. ; Gemignani, F. ; Försti, A. ; Hemminki, K. ; Elisei, R. ; Romei, C. ; Zizzi, E. A. ; Deriu, M. A. ; Simoni, M. ; Landi, S. ; Casarini, L.
Tietze, A. ; Mankad, K. ; Lequin, M. H. ; Ivarsson, L. ; Mirsky, D. ; Jaju, A. ; Kool, M. ; Hoff, K. V. ; Bison, B. ; Löbel, U.
Blümel, L. ; Qin, N. ; Berlandi, J. ; Paisana, E. ; Cascão, R. ; Custódia, C. ; Pauck, D. ; Picard, D. ; Langini, M. ; Stühler, K. ; Meyer, F.-D. ; Göbbels, S. ; Malzkorn, B. ; Liebau, M. C. ; Barata, J. T. ; Jeibmann, A. ; Kerl, K. ; Erkek, S. ; Kool, M. ; Pfister, S. M. ; Johann, P. D. ; Frühwald, M. C. ; Borkhardt, A. ; Reifenberger, G. ; Faria, C. C. ; Fischer, U. ; Hasselblatt, M. ; Bartl, J. ; Remke, M.
Sievers, P. ; Sill, M. ; Schrimpf, D. ; Friedel, D. ; Sturm, D. ; Gardberg, M. ; Kurian, K. M. ; Krskova, L. ; Vicha, A. ; Schaller, T. ; Hagel, C. ; Abdullaev, Z. ; Aldape, K. ; Jacques, T. S. ; Korshunov, A. ; Wick, W. ; Pfister, S. ; von Deimling, A. ; Jones, D. ; Sahm, F.
Reinhardt, A. ; Pfister, K. ; Schrimpf, D. ; Stichel, D. ; Sahm, F. ; Reuss, D. E. ; Capper, D. ; Wefers, A. ; Ebrahimi, A. ; Sill, M. ; Felsberg, J. ; Reifenberger, G. ; Becker, A. ; Prinz, M. ; Staszewski, O. ; Hartmann, C. ; Schittenhelm, J. ; Gramatzki, D. ; Weller, M. ; Olar, A. ; Rushing, E. J. ; Bergmann, M. ; Farrell, M. A. ; Blümcke, I. ; Coras, R. ; Beckervordersandforth, J. ; Kim, S. H. ; Rogerio, F. ; Dimova, P. S. ; Niehusmann, P. ; Unterberg, A. ; Platten, M. ; Pfister, S. ; Wick, W. ; Herold-Mende, C. ; von Deimling, A.
Penkert, J. ; Strüwe, F. J. ; Dutzmann, C. M. ; Doergeloh, B. B. ; Montellier, E. ; Freycon, C. ; Keymling, M. ; Schlemmer, H.-P. ; Sänger, B. ; Hoffmann, B. ; Gerasimov, T. ; Blattmann, C. ; Fetscher, S. ; Frühwald, M. ; Hettmer, S. ; Kordes, U. ; Ridola, V. ; Kroiss Benninger, S. ; Mastronuzzi, A. ; Schott, S. ; Nees, J. ; Prokop, A. ; Redlich, A. ; Seidel, M. G. ; Zimmermann, S. ; Pajtler, K. ; Pfister, S. ; Hainaut, P. ; Kratz, C. P.
Franceschi, E. ; Giannini, C. ; Furtner, J. ; Pajtler, K. ; Asioli, S. ; Guzman, R. ; Seidel, C. ; Gatto, L. ; Hau, P.
Bartl, J. ; Zanini, M. ; Bernardi, F. ; Forget, A. ; Blümel, L. ; Talbot, J. ; Picard, D. ; Qin, N. ; Cancila, G. ; Gao, Q. ; Nath, S. ; Koumba, I. M. ; Wolter, M. ; Kuonen, F. ; Langini, M. ; Beez, T. ; Munoz, C. ; Pauck, D. ; Marquardt, V. ; Yu, H. ; Souphron, J. ; Korsch, M. ; Mölders, C. ; Berger, D. ; Göbbels, S. ; Meyer, F.-D. ; Scheffler, B. ; Rotblat, B. ; Diederichs, S. ; Ramaswamy, V. ; Suzuki, H. ; Oro, A. ; Stühler, K. ; Stefanski, A. ; Fischer, U. ; Leprivier, G. ; Willbold, D. ; Steger, G. ; Buell, A. ; Kool, M. ; Lichter, P. ; Pfister, S. ; Northcott, P. A. ; Taylor, M. D. ; Borkhardt, A. ; Reifenberger, G. ; Ayrault, O. ; Remke, M.
Tsai, J. W. ; Cejas, P. ; Wang, D. K. ; Patel, S. ; Wu, D. W. ; Arounleut, P. ; Wei, X. ; Zhou, N. ; Syamala, S. ; Dubois, F. P. B. ; Crane, A. ; Pelton, K. ; Vogelzang, J. ; Sousa, C. ; Baguette, A. ; Chen, X. ; Condurat, A. L. ; Dixon-Clarke, S. E. ; Zhou, K. N. ; Lu, S. D. ; Gonzalez, E. M. ; Chacon, M. S. ; Digiacomo, J. J. ; Kumbhani, R. ; Novikov, D. ; Hunter, J. ; Tsoli, M. ; Ziegler, D. S. ; Dirksen, U. ; Jäger, N. ; Balasubramanian, G. ; Kramm, C. M. ; Nathrath, M. ; Bielack, S. ; Baker, S. J. ; Zhang, J. ; McFarland, J. M. ; Getz, G. ; Aguet, F. ; Jabado, N. ; Witt, O. ; Pfister, S. ; Ligon, K. L. ; Hovestadt, V. ; Kleinman, C. L. ; Long, H. ; Jones, D. ; Bandopadhayay, P. ; Phoenix, T. N.
Guerrini-Rousseau, L. ; Masliah-Planchon, J. ; Waszak, S. M. ; Alhopuro, P. ; Benusiglio, P. R. ; Bourdeaut, F. ; Brecht, I. B. ; Del Baldo, G. ; Dhanda, S. K. ; Garrè, M. L. ; Gidding, C. E. M. ; Hirsch, S. ; Hoarau, P. ; Jorgensen, M. ; Kratz, C. ; Lafay-Cousin, L. ; Mastronuzzi, A. ; Pastorino, L. ; Pfister, S. M. ; Schroeder, C. ; Smith, M. J. ; Vahteristo, P. ; Vibert, R. ; Vilain, C. ; Waespe, N. ; Winship, I. M. ; Evans, D. G. ; Brugieres, L.
Catanzaro, G. ; Besharat, Z. M. ; Carai, A. ; Jäger, N. ; Splendiani, E. ; Colin, C. ; Po, A. ; Chiacchiarini, M. ; Citarella, A. ; Gianno, F. ; Cacchione, A. ; Miele, E. ; Diomedi Camassei, F. ; Gessi, M. ; Massimi, L. ; Locatelli, F. ; Jones, D. ; Figarella-Branger, D. ; Pfister, S. ; Mastronuzzi, A. ; Giangaspero, F. ; Ferretti, E.
Richter-Pechańska, P. ; Kunz, J. B. ; Rausch, T. ; Erarslan-Uysal, B. ; Bornhauser, B. ; Frismantas, V. ; Assenov, Y. ; Zimmermann, M. ; Happich, M. ; von Knebel-Doeberitz, C. ; von Neuhoff, N. ; Köhler, R. ; Stanulla, M. ; Schrappe, M. ; Cario, G. ; Escherich, G. ; Kirschner-Schwabe, R. ; Eckert, C. ; Avigad, S. ; Pfister, S. M. ; Muckenthaler, M. U. ; Bourquin, J.-P. ; Korbel, J. O. ; Kulozik, A.
García-Martín, P. ; Díez, A. M. ; Maldonado, J. M. S. ; Serrano, A. J. C. ; Ter Horst, R. ; Benavente, Y. ; Landi, S. ; Macauda, A. ; Clay-Gilmour, A. ; Hernández-Mohedo, F. ; Niazi, Y. ; González-Sierra, P. ; Espinet, B. ; Rodríguez-Sevilla, J. J. ; Maffei, R. ; Blanco, G. ; Giaccherini, M. ; Puiggros, A. ; Cerhan, J. ; Marasca, R. ; Cañadas-Garre, M. ; López-Nevot, M. Á. ; Chen-Liang, T. ; Thomsen, H. ; Gámez, I. ; Moreno, V. ; Marcos-Gragera, R. ; García-Álvarez, M. ; Llorca, J. ; Jerez, A. ; Berndt, S. ; Butrym, A. ; Norman, A. D. ; Casabonne, D. ; Luppi, M. ; Slager, S. L. ; Hemminki, K. ; Li, Y. ; Alcoceba, M. ; Campa, D. ; Canzian, F. ; de Sanjosé, S. ; Försti, A. ; Netea, M. G. ; Jurado, M. ; Sainz, J.
Schedel, A. ; Friedrich, U. A. ; Morcos, M. N. F. ; Wagener, R. ; Mehtonen, J. ; Watrin, T. ; Saitta, C. ; Brozou, T. ; Michler, P. ; Walter, C. ; Försti, A. ; Baksi, A. ; Menzel, M. ; Horak, P. ; Paramasivam, N. ; Fazio, G. ; Autry, R. ; Fröhling, S. ; Suttorp, M. ; Gertzen, C. ; Gohlke, H. ; Bhatia, S. ; Wadt, K. ; Schmiegelow, K. ; Dugas, M. ; Richter, D. ; Glimm, H. ; Heinäniemi, M. ; Jessberger, R. ; Cazzaniga, G. ; Borkhardt, A. ; Hauer, J. ; Auer, F.
A synergistic interaction between HDAC- and PARP inhibitors in childhood tumors with chromothripsis.
Khalid, U. ; Simovic, M. ; Hammann, L. A. ; Iskar, M. ; Wong, J. K. L. ; Kumar, R. ; Jugold, M. ; Sill, M. ; Bolkestein, M. ; Kolb, T. ; Hergt, M. ; Devens, F. ; Ecker, J. ; Kool, M. ; Milde, T. ; Westermann, F. ; Benner, A. ; Lewis, J. ; Dietrich, S. ; Pfister, S. M. ; Lichter, P. ; Zapatka, M. ; Ernst, A.
Nemes, K. ; Johann, P. D. ; Steinbügl, M. ; Gruhle, M. ; Bens, S. ; Kachanov, D. ; Teleshova, M. ; Hauser, P. ; Simon, T. ; Tippelt, S. ; Eberl, W. ; Chada, M. ; Lopez, V. S. ; Grigull, L. ; Hernáiz-Driever, P. ; Eyrich, M. ; Pears, J. ; Milde, T. ; Reinhard, H. ; Leipold, A. ; van de Wetering, M. ; Gil-da-Costa, M. J. ; Ebetsberger-Dachs, G. ; Kerl, K. ; Lemmer, A. ; Boztug, H. ; Furtwängler, R. ; Kordes, U. ; Vokuhl, C. ; Hasselblatt, M. ; Bison, B. ; Kröncke, T. ; Melchior, P. ; Timmermann, B. ; Gerss, J. ; Siebert, R. ; Frühwald, M. C.
Federico, A. ; Thomas, C. ; Miskiewicz, K. ; Woltering, N. ; Zin, F. ; Nemes, K. ; Bison, B. ; Johann, P. D. ; Hawes, D. ; Bens, S. ; Kordes, U. ; Albrecht, S. ; Dohmen, H. ; Hauser, P. ; Keyvani, K. ; van Landeghem, F. K. H. ; Lund, E. L. ; Scheie, D. ; Mawrin, C. ; Monoranu, C.-M. ; Parm Ulhøi, B. ; Pietsch, T. ; Reinhard, H. ; Riemenschneider, M. J. ; Sehested, A. ; Sumerauer, D. ; Siebert, R. ; Paulus, W. ; Frühwald, M. C. ; Kool, M. ; Hasselblatt, M.
Beck, P. ; Selle, B. ; Madenach, L. ; Jones, D. T. W. ; Vokuhl, C. ; Gopisetty, A. ; Nabbi, A. ; Brecht, I. B. ; Ebinger, M. ; Wegert, J. ; Graf, N. ; Gessler, M. ; Pfister, S. M. ; Jäger, N.
Hasselblatt, M. ; Thomas, C. ; Federico, A. ; Nemes, K. ; Johann, P. D. ; Bison, B. ; Bens, S. ; Dahlum, S. ; Kordes, U. ; Redlich, A. ; Lessel, L. ; Pajtler, K. W. ; Mawrin, C. ; Schüller, U. ; Nolte, K. ; Kramm, C. M. ; Hinz, F. ; Sahm, F. ; Giannini, C. ; Penkert, J. ; Kratz, C. P. ; Pfister, S. M. ; Siebert, R. ; Paulus, W. ; Kool, M. ; Frühwald, M. C.
Clay-Gilmour, A. ; Chattopadhyay, S. ; Hildebrandt, M. A. T. ; Thomsen, H. ; Weinhold, N. ; Vodicka, P. ; Vodickova, L. ; Hoffmann, P. ; Nöthen, M. M. ; Jöckel, K.-H. ; Schmidt, B. ; Langer, C. ; Hajek, R. ; Hallmans, G. ; Pettersson-Kymmer, U. ; Ohlsson, C. ; Späth, F. ; Houlston, R. ; Goldschmidt, H. ; Manasanch, E. E. ; Norman, A. ; Kumar, S. ; Rajkumar, S. V. ; Slager, S. ; Försti, A. ; Vachon, C. M. ; Hemminki, K.
Patel Jamilkhan, A. ; Dogan, H. ; Payne, A. ; Krause, E. ; Sievers, P. ; Schoebe, N. ; Schrimpf, D. ; Blume, C. ; Stichel, D. ; Holmes, N. ; Euskirchen, P. ; Hench, J. ; Frank, S. ; Rosenstiel-Goidts, V. ; Ratliff, M. ; Etminan, N. ; Unterberg, A. ; Dieterich, C. ; Herold-Mende, C. ; Pfister, S. M. ; Wick, W. ; Loose, M. ; von Deimling, A. ; Sill, M. ; Jones, D. T. W. ; Schlesner, M. ; Sahm, F.
Graf, M. ; Interlandi, M. ; Moreno, N. ; Holdhof, D. ; Göbel, C. ; Melcher, V. ; Mertins, J. ; Albert, T. K. ; Kastrati, D. ; Alfert, A. ; Holsten, T. ; de Faria, F. ; Meisterernst, M. ; Rossig, C. ; Warmuth-Metz, M. ; Nowak, J. ; Meyer Zu Hörste, G. ; Mayère, C. ; Nef, S. ; Johann, P. ; Frühwald, M. C. ; Dugas, M. ; Schüller, U. ; Kerl, K.
Kaltenbacher, T. ; Löprich, J. ; Maresch, R. ; Weber, J. ; Müller, S. ; Oellinger, R. ; Groß, N. ; Griger, J. ; de Andrade Krätzig, N. ; Avramopoulos, P. ; Ramanujam, D. ; Brummer, S. ; Widholz, S. A. ; Bärthel, S. ; Falcomatà, C. ; Pfaus, A. ; Alnatsha, A. ; Mayerle, J. ; Schmidt-Supprian, M. ; Reichert, M. ; Schneider, G. ; Ehmer, U. ; Braun, C. J. ; Saur, D. K. M. ; Engelhardt, S. ; Rad, R.
Sun, Y.-C. ; Sun, P. ; Xue, J. ; Du, Y. ; Yan, H. ; Wang, L.-W. ; Yi, X.-X. ; Sun, J.-G. ; Zhang, X. ; Gao, J.-L.
Roosen, M. ; Odé, Z. ; Bunt, J. ; Kool, M.
Gatzweiler, C. ; Ridinger, J. ; Herter, S. ; Gerloff, X. F. ; ElHarouni, D. ; Berker, Y. ; Imle, R. ; Schmitt, L. ; Kreth, S. ; Stainczyk, S. ; Ayhan, S. ; Najafi, S. ; Krunic, D. ; Frese, K. ; Meder, B. ; Reuss, D. ; Fiesel, P. ; Schramm, K. ; Blattner-Johnson, M. ; Jones, D. T. W. ; Banito, A. ; Westermann, F. ; Oppermann, S. ; Milde, T. ; Peterziel, H. ; Witt, O. ; Oehme, I.
Gaab, C. ; Adolph, J. E. ; Tippelt, S. ; Mikasch, R. ; Obrecht, D. ; Mynarek, M. ; Rutkowski, S. ; Pfister, S. M. ; Milde, T. ; Witt, O. ; Bison, B. ; Warmuth-Metz, M. ; Kortmann, R.-D. ; Dietzsch, S. ; Pietsch, T. ; Timmermann, B. ; Sträter, R. ; Bode, U. ; Faldum, A. ; Kwiecien, R. ; Fleischhack, G.
Obrecht, D. ; Mynarek, M. ; Hagel, C. ; Kwiecien, R. ; Spohn, M. ; Bockmayr, M. ; Bison, B. ; Pfister, S. M. ; Jones, D. T. W. ; Sturm, D. ; von Deimling, A. ; Sahm, F. ; von Hoff, K. ; Juhnke, B.-O. ; Benesch, M. ; Gerber, N. U. ; Friedrich, C. ; von Bueren, A. O. ; Kortmann, R.-D. ; Schwarz, R. ; Pietsch, T. ; Fleischhack, G. ; Schüller, U. ; Rutkowski, S.
Hasselblatt, M. ; Thomas, C. ; Federico, A. ; Bens, S. ; Hellström, M. ; Casar-Borota, O. ; Kordes, U. ; Neumann, J. E. ; Dottermusch, M. ; Rodriguez, F. J. ; Lo, A. C. ; Cheng, S. ; Hendson, G. ; Hukin, J. ; Hartmann, C. ; Koch, A. ; Capper, D. ; Siebert, R. ; Paulus, W. ; Nemes, K. ; Johann, P. D. ; Frühwald, M. C. ; Kool, M.
Sun, Y.-C. ; Sun, P. ; Xue, J. ; Du, Y. ; Yan, H. ; Wang, L.-W. ; Yi, X.-X. ; Sun, J.-G. ; Zhang, X. ; Gao, J.-L.
Ebrahimi, A. ; Korshunov, A. ; Reifenberger, G. ; Capper, D. ; Felsberg, J. ; Trisolini, E. ; Pollo, B. ; Calatozzolo, C. ; Prinz, M. ; Staszewski, O. ; Schweizer, L. ; Schittenhelm, J. ; Harter, P. ; Paulus, W. ; Thomas, C. ; Kohlhof-Meinecke, P. ; Seiz-Rosenhagen, M. ; Milde, T. ; Casalini, M. B. ; Suwala, A. ; Wefers, A. K. ; Reinhardt, A. ; Sievers, P. ; Kramm, C. M. ; Etminam, N. ; Unterberg, A. ; Wick, W. ; Herold-Mende, C. ; Sturm, D. ; Pfister, S. M. ; Sill, M. ; Jones, D. T. W. ; Schrimpf, D. ; Reuss, D. E. ; Aldape, K. ; Abdullaev, Z. ; Sahm, F. ; von Deimling, A. ; Stichel, D.
Ajore, R. ; Niroula, A. ; Pertesi, M. ; Cafaro, C. ; Thodberg, M. ; Went, M. ; Bao, E. L. ; Duran-Lozano, L. ; Lopez de Lapuente Portilla, A. ; Olafsdottir, T. ; Ugidos-Damboriena, N. ; Magnusson, O. ; Samur, M. ; Lareau, C. A. ; Halldorsson, G. H. ; Thorleifsson, G. ; Norddahl, G. L. ; Gunnarsdottir, K. ; Försti, A. ; Goldschmidt, H. ; Hemminki, K. ; van Rhee, F. ; Kimber, S. ; Sperling, A. S. ; Kaiser, M. ; Anderson, K. ; Jonsdottir, I. ; Munshi, N. ; Rafnar, T. ; Waage, A. ; Weinhold, N. ; Thorsteinsdottir, U. ; Sankaran, V. G. ; Stefansson, K. ; Houlston, R. ; Nilsson, B.
Cidre-Aranaz, F. ; Li, J. ; Hölting, T. L. B. ; Orth, M. F. ; Imle, R. ; Kutschmann, S. ; Ammirati, G. ; Ceranski, K. ; Carreño-Gonzalez, M. J. ; Kasan, M. ; Marchetto, A. ; Funk, C. M. ; Bestvater, F. ; Bersini, S. ; Arrigoni, C. ; Moretti, M. ; Thiel, U. ; Baumhoer, D. ; Sahm, F. ; Pfister, S. M. ; Hartmann, W. ; Dirksen, U. ; Romero-Pérez, L. ; Banito, A. ; Ohmura, S. ; Musa, J. ; Kirchner, T. ; Knott, M. M. L. ; Grünewald, T.
Diaz Coronado, R. Y. ; Mynarek, M. ; Koelsche, C. ; Mora Alferez, P. ; Casavilca Zambrano, S. ; Wachtel Aptowitzer, A. ; Sahm, F. ; Deimling, A. ; Schüller, U. ; Spohn, M. ; Sturm, D. ; Pfister, S. M. ; Morales La Madrid, A. ; Sernaque Quintana, R. ; Sarria Bardales, G. ; Negreiros Chinchihuara, T. ; Ojeda Medina, L. ; Garcia-Corrochano Medina, P. ; Campos Sanchez, D. A. ; Ponce Farfan, J. ; Rutkowski, S. ; Garcia Leon, J. L.
Massimino, M. ; Barretta, F. ; Modena, P. ; Johann, P. ; Ferroli, P. ; Antonelli, M. ; Gandola, L. ; Garrè, M. L. ; Bertin, D. ; Mastronuzzi, A. ; Mascarin, M. ; Quaglietta, L. ; Viscardi, E. ; Sardi, I. ; Ruggiero, A. ; Boschetti, L. ; Giagnacovo, M. ; Biassoni, V. ; Schiavello, E. ; Chiapparini, L. ; Erbetta, A. ; Mussano, A. ; Giussani, C. ; Mura, R. M. ; Barra, S. ; Scarzello, G. ; Scimone, G. ; Carai, A. ; Giangaspero, F. ; Buttarelli, F. R.
Richardson, S. ; Hill, R. M. ; Kui, C. ; Lindsey, J. C. ; Grabovksa, Y. ; Keeling, C. ; Pease, L. ; Bashton, M. ; Crosier, S. ; Vinci, M. ; André, N. ; Figarella-Branger, D. ; Hansford, J. R. ; Lastowska, M. ; Zakrzewski, K. ; Jorgensen, M. ; Pickles, J. C. ; Taylor, M. D. ; Pfister, S. M. ; Wharton, S. B. ; Pizer, B. ; Michalski, A. ; Joshi, A. ; Jacques, T. S. ; Hicks, D. ; Schwalbe, E. C. ; Williamson, D. ; Ramaswamy, V. ; Bailey, S. ; Clifford, S. C.
Riemondy, K. A. ; Venkataraman, S. ; Willard, N. ; Nellan, A. ; Sanford, B. ; Griesinger, A. M. ; Amani, V. ; Mitra, S. ; Hankinson, T. C. ; Handler, M. H. ; Sill, M. ; Ocasio, J. ; Weir, S. J. ; Malawsky, D. S. ; Gershon, T. R. ; Garancher, A. ; Wechsler-Reya, R. J. ; Hesselberth, J. R. ; Foreman, N. K. ; Donson, A. M. ; Vibhakar, R.
Blume, C. ; Dogan, H. ; Schweizer, L. ; Peyre, M. ; Doll, S. ; Picard, D. ; Sankowski, R. ; Hovestadt, V. ; Okonechnikov, K. ; Sievers, P. ; Patel, A. ; Reuss, D. ; Friedrich, M. ; Stichel, D. ; Schrimpf, D. ; Beck, K. ; Wirsching, H.-G. ; Jungwirth, G. ; Hanemann, C. O. ; Lamszus, K. ; Westphal, M. ; Etminan, N. ; Unterberg, A. ; Mawrin, C. ; Remke, M. ; Ayrault, O. ; Lichter, P. ; Pfister, S. M. ; Reifenberger, G. ; Platten, M. ; Milde, T. ; Jones, D. T. ; Grossmann, R. ; Ram, Z. ; Ratliff, M. ; Herold-Mende, C. ; Mallm, J.-P. ; Neidert, M. C. ; Wick, W. ; Prinz, M. ; Weller, M. ; Mann, M. ; Kalamarides, M. ; von Deimling, A. ; Schlesner, M. ; Sahm, F.
Voellger, B. ; Zhang, Z. ; Benzel, J. ; Wang, J. ; Lei, T. ; Nimsky, C. ; Bartsch, J.-W.
Williamson, L. M. ; Rive, C. M. ; Di Francesco, D. ; Titmuss, E. ; Chun, H. E. ; Brown, S. D. ; Milne, K. ; Pleasance, E. ; Lee, A. F. ; Yip, S. ; Rosenbaum, D. G. ; Hasselblatt, M. ; Johann, P. D. ; Kool, M. ; Harvey, M. ; Dix, D. ; Renouf, D. J. ; Holt, R. A. ; Nelson, B. H. ; Hirst, M. ; Jones, S. J. M. ; Laskin, J. ; Rassekh, S. R. ; Deyell, R. J. ; Marra, M. A.
Shatara, M. ; Schieffer, K. M. ; Klawinski, D. ; Thomas, D. L. ; Pierson, C. R. ; Sribnick, E. A. ; Jones, J. ; Rodriguez, D. P. ; Deeg, C. ; Hamelberg, E. ; LaHaye, S. ; Miller, K. E. ; Fitch, J. ; Kelly, B. ; Leraas, K. ; Pfau, R. ; White, P. ; Magrini, V. ; Wilson, R. K. ; Mardis, E. R. ; Abdelbaki, M. S. ; Finlay, J. L. ; Boué, D. R. ; Cottrell, C. E. ; Ghasemi, D. R. ; Pajtler, K. W. ; Osorio, D. S.
Duran-Lozano, L. ; Thorleifsson, G. ; Lopez de Lapuente Portilla, A. ; Niroula, A. ; Went, M. ; Thodberg, M. ; Pertesi, M. ; Ajore, R. ; Cafaro, C. ; Olason, P. I. ; Stefansdottir, L. ; Bragi Walters, G. ; Halldorsson, G. H. ; Turesson, I. ; Kaiser, M. F. ; Försti, A. ; Weinhold, N. ; Abildgaard, N. ; Andersen, N. F. ; Mellqvist, U.-H. ; Waage, A. ; Juul-Vangsted, A. ; Thorsteinsdottir, U. ; Hansson, M. ; Houlston, R. ; Rafnar, T. ; Stefansson, K. ; Nilsson, B.
Kaur, A. ; Chopra, M. ; Bhushan, M. ; Gupta, S. ; Kumari P, H. ; Sivagurunathan, N. ; Shukla, N. ; Rajagopal, S. ; Bhalothia, P. ; Sharma, P. ; Naravula, J. ; Suravajhala, R. ; Gupta, A. ; Abbasi, B. A. ; Goswami, P. ; Singh, H. ; Narang, R. ; Polavarapu, R. ; Medicherla, K. M. ; Valadi, J. ; Kumar S, A. ; Chaubey, G. ; Singh, K. K. ; Bandapalli, O. R. ; Kavi Kishor, P. B. ; Suravajhala, P.
Huang, Y. ; Qi, L. ; Kogiso, M. ; Du, Y. ; Braun, F. K. ; Zhang, H. ; Huang, L. F. ; Xiao, S. ; Teo, W.-Y. ; Lindsay, H. ; Zhao, S. ; Baxter, P. ; Su, J. M. F. ; Adesina, A. ; Yang, J. ; Brabetz, S. ; Kool, M. ; Pfister, S. M. ; Chintagumpala, M. ; Perlaky, L. ; Wang, Z. ; Zhou, Y. ; Man, T.-K. ; Li, X.-N.
Liu, A. P. Y. ; Smith, K. S. ; Kumar, R. ; Paul, L. ; Bihannic, L. ; Lin, T. ; Maass, K. K. ; Pajtler, K. W. ; Chintagumpala, M. ; Su, J. M. ; Bouffet, E. ; Fisher, M. J. ; Gururangan, S. ; Cohn, R. ; Hassall, T. ; Hansford, J. R. ; Klimo, P. ; Boop, F. A. ; Stewart, C. F. ; Harreld, J. H. ; Merchant, T. E. ; Tatevossian, R. G. ; Neale, G. ; Lear, M. ; Klco, J. M. ; Orr, B. A. ; Ellison, D. W. ; Gilbertson, R. J. ; Onar-Thomas, A. ; Gajjar, A. ; Robinson, G. W. ; Northcott, P. A.
Schmid, S. ; Solomon, D. A. ; Perez, E. ; Thieme, A. ; Kleinschmidt-DeMasters, B. K. ; Giannini, C. ; Reinhardt, A. ; Asa, S. L. ; Mete, O. ; Stichel, D. ; Siewert, C. ; Dittmayer, C. ; Hasselblatt, M. ; Paulus, W. ; Nagel, C. ; Harter, P. ; Schittenhelm, J. ; Honegger, J. ; Rushing, E. ; Coras, R. ; Pfister, S. M. ; Buslei, R. ; Koch, A. ; Perry, A. ; Jones, D. T. W. ; von Deimling, A. ; Capper, D. ; Lopes, M. B.
Sievers, P. ; Stichel, D. ; Sill, M. ; Schrimpf, D. ; Sturm, D. ; Selt, F. ; Ecker, J. ; Kazdal, D. ; Miele, E. ; Kranendonk, M. E. G. ; Tops, B. B. J. ; Kohlhof-Meinecke, P. ; Beschorner, R. ; Kramm, C. M. ; Hasselblatt, M. ; Reifenberger, G. ; Capper, D. ; Wesseling, P. ; Stenzinger, A. ; Milde, T. ; Korshunov, A. ; Witt, O. ; Pfister, S. M. ; Wick, W. ; von Deimling, A. ; Jones, D. T. W. ; Sahm, F.
Deng, M. Y. ; Sturm, D. ; Pfaff, E. ; Sill, M. ; Stichel, D. ; Balasubramanian, G. P. ; Tippelt, S. ; Kramm, C. ; Donson, A. M. ; Green, A. L. ; Jones, C. ; Schittenhelm, J. ; Ebinger, M. ; Schuhmann, M. U. ; Jones, B. C. ; van Tilburg, C. M. ; Wittmann, A. ; Golanov, A. ; Ryzhova, M. ; Ecker, J. ; Milde, T. ; Witt, O. ; Sahm, F. ; Reuss, D. ; Sumerauer, D. ; Zamecnik, J. ; Korshunov, A. ; von Deimling, A. ; Pfister, S. M. ; Jones, D.
Marcu, A. ; Schlosser, A. ; Keupp, A. ; Trautwein, N. ; Johann, P. ; Wölfl, M. ; Lager, J. ; Monoranu, C. M. ; Walz, J. S. ; Henkel, L. M. ; Krauß, J. ; Ebinger, M. ; Schuhmann, M. ; Thomale, U. W. ; Pietsch, T. ; Klinker, E. ; Schlegel, P. G. ; Oyen, F. ; Reisner, Y. ; Rammensee, H.-G. ; Eyrich, M.
Panwalkar, P. ; Tamrazi, B. ; Dang, D. ; Chung, C. ; Sweha, S. ; Natarajan, S. K. ; Pun, M. ; Bayliss, J. ; Ogrodzinski, M. P. ; Pratt, D. ; Mullan, B. ; Hawes, D. ; Yang, F. ; Lu, C. ; Sabari, B. R. ; Achreja, A. ; Heon, J. ; Animasahun, O. ; Cieslik, M. ; Dunham, C. ; Yip, S. ; Hukin, J. ; Phillips, J. J. ; Bornhorst, M. ; Griesinger, A. M. ; Donson, A. M. ; Foreman, N. K. ; Garton, H. J. L. ; Heth, J. ; Muraszko, K. ; Nazarian, J. ; Koschmann, C. ; Jiang, L. ; Filbin, M. G. ; Nagrath, D. ; Kool, M. ; Korshunov, A. ; Pfister, S. ; Gilbertson, R. J. ; Allis, C. D. ; Chinnaiyan, A. M. ; Lunt, S. Y. ; Blüml, S. ; Judkins, A. R. ; Venneti, S.
Kirches, E. ; Sahm, F. ; Korshunov, A. ; Bluecher, C. ; Waldt, N. ; Kropf, S. ; Schrimpf, D. ; Sievers, P. ; Stichel, D. ; Schüller, U. ; Schittenhelm, J. ; Riemenschneider, M. J. ; Karajannis, M. A. ; Perry, A. ; Pietsch, T. ; Boekhoff, S. ; Capper, D. ; Beck, K. ; Paramasivam, N. ; Schlesner, M. ; Brastianos, P. K. ; Müller, H. L. ; Pfister, S. M. ; Mawrin, C.
Coltin, H. ; Sundaresan, L. ; Smith, K. S. ; Skowron, P. ; Massimi, L. ; Eberhart, C. G. ; Schreck, K. C. ; Gupta, N. ; Weiss, W. A. ; Tirapelli, D. ; Carlotti, C. ; Li, K. K. W. ; Ryzhova, M. ; Golanov, A. ; Zheludkova, O. ; Absalyamova, O. ; Okonechnikov, K. ; Stichel, D. ; von Deimling, A. ; Giannini, C. ; Raskin, S. ; Van Meir, E. G. ; Chan, J. A. ; Fults, D. ; Chambless, L. B. ; Kim, S.-K. ; Vasiljevic, A. ; Faure-Conter, C. ; Vibhakar, R. ; Jung, S. ; Leary, S. ; Mora, J. ; McLendon, R. E. ; Pollack, I. F. ; Hauser, P. ; Grajkowska, W. A. ; Rubin, J. B. ; van Veelen, M. C. ; French, P. J. ; Kros, J. M. ; Liau, L. M. ; Pfister, S. M. ; Kool, M. ; Kijima, N. ; Taylor, M. D. ; Packer, R. J. ; Northcott, P. A. ; Korshunov, A. ; Ramaswamy, V.
Steinbügl, M. ; Nemes, K. ; Johann, P. ; Kröncke, T. ; Tüchert, S. ; da Costa, M. J. G. ; Ebinger, M. ; Schüller, U. ; Sehested, A. ; Hauser, P. ; Reinhard, H. ; Sumerauer, D. ; Hettmer, S. ; Jakob, M. ; Hasselblatt, M. ; Siebert, R. ; Witt, O. ; Gerss, J. ; Kerl, K. ; Frühwald, M. C.
Koelsche, C. ; Benhamida, J. K. ; Kommoss, F. K. F. ; Stichel, D. ; Jones, D. ; Pfister, S. M. ; Heilig, C. E. ; Fröhling, S. ; Stenzinger, A. ; Buslei, R. ; Mentzel, T. ; Baumhoer, D. ; Ladanyi, M. ; Antonescu, C. R. ; Flucke, U. ; Gorp, J. v. ; Bode-Lesniewska, B. ; von Deimling, A. ; Mechtersheimer, G.
Hau, P. ; Frappaz, D. ; Hovey, E. ; McCabe, M. G. ; Pajtler, K. W. ; Wiestler, B. ; Seidel, C. ; Combs, S. E. ; Dirven, L. ; Klein, M. ; Anazodo, A. ; Hattingen, E. ; Hofer, S. ; Pfister, S. M. ; Zimmer, C. ; Kortmann, R.-D. ; Sunyach, M.-P. ; Tanguy, R. ; Effeney, R. ; von Deimling, A. ; Sahm, F. ; Rutkowski, S. ; Berghoff, A. S. ; Franceschi, E. ; Pineda, E. ; Beier, D. ; Peeters, E. ; Gorlia, T. ; Vanlancker, M. ; Bromberg, J. E. C. ; Gautier, J. ; Ziegler, D. S. ; Preusser, M. ; Wick, W. ; Weller, M.
Glöss, S. ; Jurmeister, P. ; Thieme, A. ; Schmid, S. ; Cai, W. Y. ; Serrette, R. N. ; Perner, S. ; Ribbat-Idel, J. ; Pagenstecher, A. ; Bläker, H. ; Keber, U. ; Stadelmann, C. ; Zechel, S. ; Johann, P. D. ; Hasselblatt, M. ; Paulus, W. ; Thomas, C. ; Dohmen, H. ; Baumhoer, D. ; Frank, S. ; Agaimy, A. ; Schüller, U. ; Vasudevaraja, V. ; Snuderl, M. ; Liu, C. Z. ; Pfister, D. G. ; Jungbluth, A. A. ; Ghossein, R. A. ; Xu, B. ; Capper, D. ; Dogan, S.
Saunders, C. N. ; Chattopadhyay, S. ; Huhn, S. ; Weinhold, N. ; Hoffmann, P. ; Nöthen, M. M. ; Jöckel, K.-H. ; Schmidt, B. ; Landi, S. ; Goldschmidt, H. ; Milani, P. ; Merlini, G. ; Rowcieno, D. ; Hawkins, P. ; Hegenbart, U. ; Palladini, G. ; Wechalekar, A. ; Schönland, S. O. ; Försti, A. ; Houlston, R. ; Hemminki, K.
Young, M. D. ; Mitchell, T. J. ; Custers, L. ; Margaritis, T. ; Morales-Rodriguez, F. ; Kwakwa, K. ; Khabirova, E. ; Kildisiute, G. ; Oliver, T. R. W. ; de Krijger, R. R. ; van den Heuvel-Eibrink, M. M. ; Comitani, F. ; Piapi, A. ; Bugallo-Blanco, E. ; Thevanesan, C. ; Burke, C. ; Prigmore, E. ; Ambridge, K. ; Roberts, K. ; Braga, F. A. V. ; Coorens, T. H. H. ; Del Valle, I. ; Wilbrey-Clark, A. ; Mamanova, L. ; Stewart, G. D. ; Gnanapragasam, V. J. ; Rampling, D. ; Sebire, N. ; Coleman, N. ; Hook, L. ; Warren, A. ; Haniffa, M. ; Kool, M. ; Pfister, S. M. ; Achermann, J. C. ; He, X. ; Barker, R. A. ; Shlien, A. ; Bayraktar, O. A. ; Teichmann, S. A. ; Holstege, F. C. ; Meyer, K. B. ; Drost, J. ; Straathof, K. ; Behjati, S.
Maass, K. ; Schad, P. ; Finster, A. ; Puranachot, P. ; Rosing, F. ; Wedig, T. ; Schwarz, N. ; Stumpf, N. ; Pfister, S. M. ; Pajtler, K.
Lötsch, D. ; Kirchhofer, D. ; Englinger, B. ; Jiang, L. ; Okonechnikov, K. ; Senfter, D. ; Laemmerer, A. ; Gabler, L. ; Pirker, C. ; Donson, A. M. ; Bannauer, P. ; Korbel, P. ; Jaunecker, C. N. ; Hübner, J.-M. ; Mayr, L. ; Madlener, S. ; Schmook, M. T. ; Ricken, G. ; Maaß, K. ; Grusch, M. ; Holzmann, K. ; Grasl-Kraupp, B. ; Spiegl-Kreinecker, S. ; Hsu, J. ; Dorfer, C. ; Rössler, K. ; Azizi, A. A. ; Foreman, N. K. ; Peyrl, A. ; Haberler, C. ; Czech, T. ; Slavc, I. ; Filbin, M. G. ; Pajtler, K. W. ; Kool, M. ; Berger, W. ; Gojo, J.
Simovic, M. ; Bolkestein, M. ; Moustafa, M. ; Wong, J. K. L. ; Körber, V. ; Benedetto, S. ; Khalid, U. ; Schreiber, H. S. ; Jugold, M. ; Korshunov, A. ; Hübschmann, D. ; Mack, N. ; Brons, S. ; Wei, P.-C. ; Breckwoldt, M. O. ; Heiland, S. ; Bendszus, M. ; Jürgen, D. ; Höfer, T. ; Zapatka, M. ; Kool, M. ; Pfister, S. M. ; Abdollahi, A. ; Ernst, A.
Gao, J.-l. ; Beck, P. ; Sun, Y.-c. ; Xue, J. ; Wang, G. ; Wang, L.-w. ; Du, Y. ; Zhang, X. ; Sun, J.-g.
Pathak, R. ; Zin, F. ; Thomas, C. ; Bens, S. ; Gayden, T. ; Karamchandani, J. ; Dudley, R. W. ; Nemes, K. ; Johann, P. D. ; Oyen, F. ; Kordes, U. ; Jabado, N. ; Siebert, R. ; Paulus, W. ; Kool, M. ; Frühwald, M. C. ; Albrecht, S. ; Kalpana, G. V. ; Hasselblatt, M.
Lyskjaer, I. ; De Noon, S. ; Tirabosco, R. ; Rocha, A. M. ; Lindsay, D. ; Amary, F. ; Ye, H. ; Schrimpf, D. ; Stichel, D. ; Sill, M. ; Koelsche, C. ; Pillay, N. ; Von Deimling, A. ; Beck, S. ; Flanagan, A. M.
Thomas, C. ; Oehl-Huber, K. ; Bens, S. ; Soschinski, P. ; Koch, A. ; Nemes, K. ; Oyen, F. ; Kordes, U. ; Kool, M. ; Frühwald, M. C. ; Hasselblatt, M. ; Siebert, R.
Zheng, T. ; Ghasemi, D. R. ; Okonechnikov, K. ; Korshunov, A. ; Sill, M. ; Maass, K. K. ; Benites Goncalves da Silva, P. ; Ryzhova, M. ; Gojo, J. ; Stichel, D. ; Arabzade, A. ; Kupp, R. ; Benzel, J. ; Taya, S. ; Adachi, T. ; Shiraishi, R. ; Gerber, N. U. ; Sturm, D. ; Ecker, J. ; Sievers, P. ; Selt, F. ; Chapman, R. ; Haberler, C. ; Figarella-Branger, D. ; Reifenberger, G. ; Fleischhack, G. ; Rutkowski, S. ; Donson, A. M. ; Ramaswamy, V. ; Capper, D. ; Ellison, D. W. ; Herold-Mende, C. C. ; Schuller, U. ; Brandner, S. ; Hernaiz Driever, P. ; Kros, J. M. ; Snuderl, M. ; Milde, T. ; Grundy, R. G. ; Hoshino, M. ; Mack, S. C. ; Gilbertson, R. J. ; Jones, D. T. W. ; Kool, M. ; von Deimling, A. ; Pfister, S. M. ; Sahm, F. ; Kawauchi, D. ; Pajtler, K.
Suwala, A. K. ; Stichel, D. ; Schrimpf, D. ; Maas, S. L. N. ; Sill, M. ; Dohmen, H. ; Banan, R. ; Reinhardt, A. ; Sievers, P. ; Hinz, F. ; Blattner-Johnson, M. ; Hartmann, C. ; Schweizer, L. ; Boldt, H. B. ; Kristensen, B. W. ; Schittenhelm, J. ; Wood, M. D. ; Chotard, G. ; Bjergvig, R. ; Das, A. ; Tabori, U. ; Hasselblatt, M. ; Korshunov, A. ; Abdullaev, Z. ; Quezado, M. ; Aldape, K. ; Harter, P. ; Snuderl, M. ; Hench, J. ; Frank, S. ; Acker, T. ; Brandner, S. ; Winkler, F. ; Wesseling, P. ; Pfister, S. M. ; Reuss, D. E. ; Wick, W. ; von Deimling, A. ; Jones, D. T. W. ; Sahm, F.
Kommoss, F. K. ; Stichel, D. ; Mora, J. ; Esteller, M. ; Jones, D. ; Pfister, S. M. ; Brack, E. ; Wachtel, M. ; Bode, P. K. ; Sinn, H.-P. ; Schmidt, D. ; Mentzel, T. ; Kommoss, F. ; Sahm, F. ; von Deimling, A. ; Koelsche, C.
ZFTA-RELA Dictates Oncogenic Transcriptional Programs to Drive Aggressive Supratentorial Ependymoma.
Arabzade, A. ; Zhao, Y. ; Varadharajan, S. ; Chen, H.-C. ; Jessa, S. ; Rivas, B. ; Stuckert, A. J. ; Solis, M. ; Kardian, A. ; Tlais, D. ; Golbourn, B. J. ; Stanton, A. J. ; Chan, Y. S. ; Olson, C. ; Karlin, K. L. ; Kong, K. ; Kupp, R. ; Hu, B. ; Injac, S. G. ; Ngo, M. ; Wang, P. R. ; De Leon, L. A. ; Sahm, F. ; Kawauchi, D. ; Pfister, S. ; Lin, C. Y. ; Hodges, H. C. ; Singh, I. ; Westbrook, T. F. ; Chintagumpala, M. M. ; Blaney, S. M. ; Parsons, D. W. ; Pajtler, K. W. ; Agnihotri, S. ; Gilbertson, R. J. ; Yi, J. ; Jabado, N. ; Kleinman, C. L. ; Bertrand, K. C. ; Deneen, B. ; Mack, S. C.
Kupp, R. ; Ruff, L. ; Terranova, S. ; Nathan, E. ; Ballereau, S. ; Stark, R. ; Sekhar Reddy Chilamakuri, C. ; Hoffmann, N. ; Wickham-Rahrmann, K. ; Widdess, M. ; Arabzade, A. ; Zhao, Y. ; Varadharajan, S. ; Zheng, T. ; Murugesan, M. K. ; Pfister, S. ; Kawauchi, D. ; Pajtler, K. W. ; Deneen, B. ; Mack, S. C. ; Masih, K. E. ; Gryder, B. E. ; Khan, J. ; Gilbertson, R. J.
Upadhyaya, S. A. ; Robinson, G. ; Onar-Thomas, A. ; Orr, B. A. ; Johann, P. ; Wu, G. ; Billups, C. A. ; Tatevossian, R. G. ; Dhanda, S. K. ; Srinivasan, A. ; Broniscer, A. ; Qaddoumi, I. ; Vinitsky, A. ; Armstrong, G. T. ; Bendel, A. ; Hassall, T. E. ; Partap, S. ; Fisher, P. G. ; Crawford, J. R. ; Chintagumpala, M. M. ; Bouffet, E. ; Gururangan, S. ; Mostafavi, R. ; Sanders, R. P. ; Klimo, P. ; Patay, Z. ; Indelicato, D. J. ; Nichols, K. E. ; Boop, F. A. ; Merchant, T. E. ; Kool, M. ; Ellison, D. W. ; Gajjar, A.
Thomas, C. ; Federico, A. ; Sill, M. ; Bens, S. ; Oyen, F. ; Nemes, K. ; Johann, P. D. ; Hartmann, C. ; Hartmann, W. ; Sumerauer, D. ; Paterno, V. ; Samii, A. ; Kordes, U. ; Siebert, R. ; Frühwald, M. C. ; Paulus, W. ; Kool, M. ; Hasselblatt, M.
Sainz, J. ; García-Verdejo, F. J. ; Martínez-Bueno, M. ; Kumar, A. ; Sánchez-Maldonado, J. M. ; Díez-Villanueva, A. ; Vodičková, L. ; Vymetálková, V. ; Martin Sánchez, V. ; Da Silva Filho, M. I. ; Sampaio-Marques, B. ; Brezina, S. ; Butterbach, K. ; ter Horst, R. ; Hoffmeister, M. ; Ludovico, P. ; Jurado, M. ; Li, Y. ; Sánchez-Rovira, P. ; Netea, M. G. ; Gsur, A. ; Vodička, P. ; Moreno, V. ; Hemminki, K. ; Brenner, H. ; Chang-Claude, J. ; Försti, A.
Custers, L. ; Khabirova, E. ; Coorens, T. H. H. ; Oliver, T. R. W. ; Calandrini, C. ; Young, M. D. ; Vieira Braga, F. A. ; Ellis, P. ; Mamanova, L. ; Segers, H. ; Maat, A. ; Kool, M. ; Hoving, E. W. ; van den Heuvel-Eibrink, M. M. ; Nicholson, J. ; Straathof, K. ; Hook, L. ; de Krijger, R. R. ; Trayers, C. ; Allinson, K. ; Behjati, S. ; Drost, J.
Hartlieb, S. ; Sieverling, L. ; Nadler-Holly, M. ; Ziehm, M. ; Toprak, U. H. ; Herrmann, C. ; Ishaque, N. ; Okonechnikov, K. ; Gartlgruber, M. ; Park, Y.-G. ; Wecht, E. M. ; Savelyeva, L. ; Henrich, K.-O. ; Rosswog, C. ; Fischer, M. ; Hero, B. ; Jones, D. T. W. ; Pfaff, E. ; Witt, O. ; Pfister, S. M. ; Volckmann, R. ; Koster, J. ; Kiesel, K. ; Rippe, K. ; Taschner-Mandl, S. ; Ambros, P. ; Brors, B. ; Selbach, M. ; Feuerbach, L. ; Westermann, F.
Liu, A. P. Y. ; Li, B. K. ; Pfaff, E. ; Gudenas, B. ; Vasiljevic, A. ; Orr, B. A. ; Dufour, C. ; Snuderl, M. ; Karajannis, M. A. ; Rosenblum, M. K. ; Hwang, E. I. ; Ng, H.-K. ; Hansford, J. R. ; Szathmari, A. ; Faure-Conter, C. ; Merchant, T. E. ; Levine, M. ; Bouvier, N. ; von Hoff, K. ; Mynarek, M. ; Rutkowski, S. ; Sahm, F. ; Kool, M. ; Hawkins, C. ; Onar-Thomas, A. ; Robinson, G. W. ; Gajjar, A. ; Pfister, S. M. ; Bouffet, E. ; Northcott, P. A. ; Jones, D. T. W. ; Huang, A.
Fisher, M. J. ; Jones, D. T. W. ; Li, Y. ; Guo, X. ; Sonawane, P. S. ; Waanders, A. J. ; Phillips, J. J. ; Weiss, W. A. ; Resnick, A. C. ; Gosline, S. ; Banerjee, J. ; Guinney, J. ; Gnekow, A. ; Kandels, D. ; Foreman, N. K. ; Korshunov, A. ; Ryzhova, M. ; Massimi, L. ; Gururangan, S. ; Kieran, M. W. ; Wang, Z. ; Fouladi, M. ; Sato, M. ; Øra, I. ; Holm, S. ; Markham, S. J. ; Beck, P. ; Jäger, N. ; Wittmann, A. ; Sommerkamp, A. C. ; Sahm, F. ; Pfister, S. ; Gutmann, D. H.
Baroni, L. ; Sundaresan, L. ; Heled, A. ; Coltin, H. ; Pajtler, K. W. ; Lin, T. ; Merchant, T. E. ; McLendon, R. ; Faria, C. ; Buntine, M. ; White, C. L. ; Pfister, S. M. ; Gilbert, M. R. ; Armstrong, T. S. ; Bouffet, E. ; Kumar, S. ; Taylor, M. D. ; Aldape, K. D. ; Ellison, D. W. ; Gottardo, N. G. ; Kool, M. ; Korshunov, A. ; Hansford, J. R. ; Ramaswamy, V.
Truong, T. ; Lesueur, F. ; Sugier, P.-E. ; Guibon, J. ; Xhaard, C. ; Karimi, M. ; Kulkarni, O. ; Lucotte, E. A. ; Bacq-Daian, D. ; Boland-Auge, A. ; Mulot, C. ; Laurent-Puig, P. ; Schvartz, C. ; Guizard, A.-V. ; Ren, Y. ; Adjadj, E. ; Rachédi, F. ; Borson-Chazot, F. ; Ortiz, R. M. ; Lence-Anta, J. J. ; Pereda, C. M. ; Comiskey, D. F. ; He, H. ; Liyanarachchi, S. ; de la Chapelle, A. ; Elisei, R. ; Gemignani, F. ; Thomsen, H. ; Forsti, A. ; Herzig, A. F. ; Leutenegger, A.-L. ; Rubino, C. ; Ostroumova, E. ; Kesminiene, A. ; Boutron-Ruault, M.-C. ; Deleuze, J.-F. ; Guénel, P. ; de Vathaire, F.
Endersby, R. ; Whitehouse, J. ; Pribnow, A. ; Kuchibhotla, M. ; Hii, H. ; Carline, B. ; Gande, S. ; Stripay, J. ; Ancliffe, M. ; Howlett, M. ; Schoep, T. ; George, C. ; Andradas, C. ; Dyer, P. ; Schluck, M. ; Patterson, B. ; Tacheva-Gigorova, S. K. ; Cooper, M. N. ; Robinson, G. ; Stewart, C. ; Pfister, S. M. ; Kool, M. ; Milde, T. ; Gajjar, A. ; Johns, T. ; Wechsler-Reya, R. J. ; Roussel, M. F. ; Gottardo, N. G.
Gajjar, A. ; Robinson, G. W. ; Smith, K. S. ; Lin, T. ; Merchant, T. E. ; Chintagumpala, M. ; Mahajan, A. ; Su, J. ; Bouffet, E. ; Bartels, U. ; Schechter, T. ; Hassall, T. ; Robertson, T. ; Nicholls, W. ; Gururangan, S. ; Schroeder, K. ; Sullivan, M. ; Wheeler, G. ; Hansford, J. R. ; Kellie, S. J. ; McCowage, G. ; Cohn, R. ; Fisher, M. J. ; Krasin, M. J. ; Stewart, C. F. ; Broniscer, A. ; Buchhalter, I. ; Tatevossian, R. G. ; Orr, B. A. ; Neale, G. ; Klimo, P. ; Boop, F. ; Srinivasan, A. ; Pfister, S. M. ; Gilbertson, R. J. ; Onar-Thomas, A. ; Ellison, D. W. ; Northcott, P. A.
Garcia-Moure, M. ; Gonzalez-Huarriz, M. ; Labiano, S. ; Guruceaga, E. ; Bandres, E. ; Zalacain, M. ; Marrodan, L. ; de Andrea, C. E. ; Villalba Esparza, M. ; Martínez-Vélez, N. ; Laspidea, V. ; Puigdelloses, M. ; Gallego Perez-Larraya, J. ; Iñigo-Marco, I. ; Stripecke, R. ; Chan, J. A. ; Raabe, E. H. ; Kool, M. ; Gomez-Manzano, C. ; Fueyo, J. ; Patiño-García, A. ; Alonso, M. M.
Adolph, J. E. ; Fleischhack, G. ; Mikasch, R. ; Zeller, J. ; Warmuth-Metz, M. ; Bison, B. ; Mynarek, M. ; Rutkowski, S. ; Schüller, U. ; von Hoff, K. ; Obrecht, D. ; Pietsch, T. ; Pfister, S. M. ; Pajtler, K. W. ; Witt, O. ; Witt, H. ; Kortmann, R.-D. ; Timmermann, B. ; Krauß, J. ; Frühwald, M. C. ; Faldum, A. ; Kwiecien, R. ; Bode, U. ; Tippelt, S. ; HIT-Network, G. G.
Holdhof, D. ; Johann, P. D. ; Spohn, M. ; Bockmayr, M. ; Safaei, S. ; Joshi, P. ; Masliah-Planchon, J. ; Ho, B. ; Andrianteranagna, M. ; Bourdeaut, F. ; Huang, A. ; Kool, M. ; Upadhyaya, S. A. ; Bendel, A. E. ; Indenbirken, D. ; Foulkes, W. D. ; Bush, J. W. ; Creytens, D. ; Kordes, U. ; Frühwald, M. C. ; Hasselblatt, M. ; Schüller, U.
Stichel, D. ; Schrimpf, D. ; Sievers, P. ; Reinhardt, A. ; Suwala, A. K. ; Sill, M. ; Reuss, D. E. ; Korshunov, A. ; Casalini, B. M. ; Sommerkamp, A. C. ; Ecker, J. ; Selt, F. ; Sturm, D. ; Gnekow, A. ; Koch, A. ; Simon, M. ; Hernáiz Driever, P. ; Schüller, U. ; Capper, D. ; van Tilburg, C. M. ; Witt, O. ; Milde, T. ; Pfister, S. M. ; Jones, D. T. W. ; von Deimling, A. ; Sahm, F. ; Wefers, A.
Sievers, P. ; Sill, M. ; Blume, C. ; Tauziede-Espariat, A. ; Schrimpf, D. ; Stichel, D. ; Reuss, D. E. ; Dogan, H. ; Hartmann, C. ; Mawrin, C. ; Hasselblatt, M. ; Stummer, W. ; Schick, U. ; Hench, J. ; Frank, S. ; Ketter, R. ; Schweizer, L. ; Schittenhelm, J. ; Puget, S. ; Brandner, S. ; Jaunmuktane, Z. ; Küsters, B. ; Abdullaev, Z. ; Pekmezci, M. ; Snuderl, M. ; Ratliff, M. ; Herold-Mende, C. ; Unterberg, A. ; Aldape, K. ; Ellison, D. W. ; Wesseling, P. ; Reifenberger, G. ; Wick, W. ; Perry, A. ; Varlet, P. ; Pfister, S. M. ; Jones, D. T. W. ; von Deimling, A. ; Sahm, F.
Gartlgruber, M. ; Sharma, A. K. ; Quintero, A. ; Dreidax, D. ; Jansky, S. ; Park, Y.-G. ; Kreth, S. ; Meder, J. ; Doncevic, D. ; Saary, P. ; Toprak, U. H. ; Ishaque, N. ; Afanasyeva, E. ; Wecht, E. ; Koster, J. ; Versteeg, R. ; Grünewald, T. ; Jones, D. T. W. ; Pfister, S. M. ; Henrich, K.-O. ; van Nes, J. ; Herrmann, C. ; Westermann, F.
Thomas, C. ; Soschinski, P. ; Zwaig, M. ; Oikonomopoulos, S. ; Okonechnikov, K. ; Pajtler, K. ; Sill, M. ; Schweizer, L. ; Koch, A. ; Neumann, J. ; Schüller, U. ; Sahm, F. ; Rauschenbach, L. ; Keyvani, K. ; Proescholdt, M. ; Riemenschneider, M. J. ; Segewiß, J. ; Ruckert, C. ; Grauer, O. ; Monoranu, C.-M. ; Lamszus, K. ; Patrizi, A. ; Kordes, U. ; Siebert, R. ; Kool, M. ; Ragoussis, J. ; Foulkes, W. D. ; Paulus, W. ; Rivera, B. ; Hasselblatt, M.
Suwala, A. K. ; Stichel, D. ; Schrimpf, D. ; Kloor, M. ; Wefers, A. K. ; Reinhardt, A. ; Maas, S. L. N. ; Kratz, C. P. ; Schweizer, L. ; Hasselblatt, M. ; Snuderl, M. ; Abedalthagafi, M. S. J. ; Abdullaev, Z. ; Monoranu, C. M. ; Bergmann, M. ; Pekrun, A. ; Freyschlag, C. ; Aronica, E. ; Kramm, C. M. ; Hinz, F. ; Sievers, P. ; Korshunov, A. ; Kool, M. ; Pfister, S. M. ; Sturm, D. ; Jones, D. T. W. ; Wick, W. ; Unterberg, A. ; Hartmann, C. ; Dodgshun, A. ; Tabori, U. ; Wesseling, P. ; Sahm, F. ; von Deimling, A. ; Reuß, D.
Albert, T. K. ; Interlandi, M. ; Sill, M. ; Graf, M. ; Moreno, N. ; Menck, K. ; Rohlmann, A. ; Melcher, V. ; Korbanka, S. ; Meyer zu Hörste, G. ; Lautwein, T. ; Frühwald, M. C. ; Krebs, C. F. ; Holdhof, D. ; Schoof, M. ; Missler, M. ; Bleckmann, A. ; Dugas, M. ; Schüller, U. ; Jäger, N. ; Pfister, S. ; Kerl, K.
Sievers, P. ; Sill, M. ; Schrimpf, D. ; Stichel, D. ; Reuß, D. ; Sturm, D. ; Hench, J. ; Frank, S. ; Krskova, L. ; Vicha, A. ; Zapotocky, M. ; Bison, B. ; Castel, D. ; Grill, J. ; Debily, M.-A. ; Harter, P. N. ; Snuderl, M. ; Kramm, C. M. ; Reifenberger, G. ; Korshunov, A. ; Jabado, N. ; Wesseling, P. ; Wick, W. ; Solomon, D. A. ; Perry, A. ; Jacques, T. S. ; Jones, C. ; Witt, O. ; Pfister, S. ; von Deimling, A. ; Jones, D. ; Sahm, F.
Massimino, M. ; Barretta, F. ; Modena, P. ; Witt, H. ; Minasi, S. ; Pfister, S. M. ; Pajtler, K. W. ; Antonelli, M. ; Gandola, L. ; Garrè, M. L. ; Bertin, D. ; Mastronuzzi, A. ; Mascarin, M. ; Quaglietta, L. ; Viscardi, E. ; Sardi, I. ; Ruggiero, A. ; Pollo, B. ; Buccoliero, A. ; Boschetti, L. ; Schiavello, E. ; Chiapparini, L. ; Erbetta, A. ; Morra, I. ; Gessi, M. ; Donofrio, V. ; Patriarca, C. ; Giangaspero, F. ; Johann, P. ; Buttarelli, F. R.
Ecker, J. ; Thatikonda, V. ; Sigismondo, G. ; Selt, F. ; Valinciute, G. ; Oehme, I. ; Müller, C. ; Buhl, J. L. ; Ridinger, J. ; Usta, D. ; Qin, N. ; van Tilburg, C. M. ; Herold-Mende, C. ; Remke, M. ; Sahm, F. ; Westermann, F. ; Kool, M. ; Wechsler-Reya, R. J. ; Chavez, L. ; Krijgsveld, J. ; Jäger, N. ; Pfister, S. M. ; Witt, O. ; Milde, T.
Kolb, T. ; Khalid, U. ; Simović, M. ; Ratnaparkhe, M. ; Wong, J. ; Jauch, A. ; Schmezer, P. ; Rode, A. ; Sebban, S. ; Haag, D. ; Hergt, M. ; Devens, F. ; Buganim, Y. ; Zapatka, M. ; Lichter, P. ; Ernst, A.
Reyna, M. A. ; Haan, D. ; Paczkowska, M. ; Verbeke, L. P. C. ; Vazquez, M. ; Kahraman, A. ; Pulido-Tamayo, S. ; Barenboim, J. ; Wadi, L. ; Dhingra, P. ; Shrestha, R. ; Getz, G. ; Lawrence, M. S. ; Pedersen, J. S. ; Rubin, M. A. ; Wheeler, D. A. ; Brunak, S. ; Izarzugaza, J. M. G. ; Khurana, E. ; Marchal, K. ; von Mering, C. ; Sahinalp, S. C. ; Valencia, A. ; Drivers, P. ; FunctionalInterpretationWorkingGroup ; Reimand, J. ; Stuart, J. M. ; Raphael, B. J. ; PCAWGConsortium
Alexandrov, L. B. ; Kim, J. ; Haradhvala, N. J. ; Huang, M. N. ; Tian Ng, A. W. ; Wu, Y. ; Boot, A. ; Covington, K. R. ; Gordenin, D. A. ; Bergstrom, E. N. ; Islam, S. M. A. ; Lopez-Bigas, N. ; Klimczak, L. J. ; McPherson, J. R. ; Morganella, S. ; Sabarinathan, R. ; Wheeler, D. A. ; Mustonen, V. ; PCAWGMutationalSignaturesWorkingGroup ; Getz, G. ; Rozen, S. G. ; Stratton, M. R. ; PCAWGConsortium
Mayr, L. ; Gojo, J. ; Peyrl, A. ; Azizi, A. A. ; Stepien, N. M. ; Pletschko, T. ; Czech, T. ; Dorfer, C. ; Lambo, S. ; Dieckmann, K. ; Haberler, C. ; Kool, M. ; Slavc, I.
Mayr, L. ; Guntner, A. S. ; Madlener, S. ; Schmook, M. T. ; Peyrl, A. ; Azizi, A. A. ; Dieckmann, K. ; Reisinger, D. ; Stepien, N. M. ; Schramm, K. ; Laemmerer, A. ; Jones, D. T. W. ; Ecker, J. ; Sahm, F. ; Milde, T. ; Pajtler, K. W. ; Blattner-Johnson, M. ; Strbac, M. ; Dorfer, C. ; Czech, T. ; Kirchhofer, D. ; Gabler, L. ; Berger, W. ; Haberler, C. ; Müllauer, L. ; Buchberger, W. ; Slavc, I. ; Lötsch-Gojo, D. ; Gojo, J.
Dietzsch, S. ; Placzek, F. ; Pietschmann, K. ; von Bueren, A. O. ; Matuschek, C. ; Glück, A. ; Guckenberger, M. ; Budach, V. ; Welzel, J. ; Pöttgen, C. ; Schmidberger, H. ; Heinzelmann, F. ; Paulsen, F. ; Escudero, M. P. ; Schwarz, R. ; Hornung, D. ; Martini, C. ; Grosu, A. L. ; Stueben, G. ; Jablonska, K. ; Dunst, J. ; Stranzl-Lawatsch, H. ; Dieckmann, K. ; Timmermann, B. ; Pietsch, T. ; Warmuth-Metz, M. ; Bison, B. ; Kwiecien, R. ; Benesch, M. ; Gerber, N. U. ; Grotzer, M. A. ; Pfister, S. M. ; Clifford, S. C. ; von Hoff, K. ; Klagges, S. ; Rutkowski, S. ; Kortmann, R.-D. ; Mynarek, M.
Rusert, J. M. ; Juarez, E. F. ; Brabetz, S. ; Jensen, J. ; Garancher, A. ; Chau, L. Q. ; Tacheva-Grigorova, S. K. ; Wahab, S. ; Udaka, Y. T. ; Finlay, D. ; Seker-Cin, H. ; Reardon, B. ; Gröbner, S. ; Serrano, J. ; Ecker, J. ; Qi, L. ; Kogiso, M. ; Du, Y. ; Baxter, P. A. ; Henderson, J. J. ; Berens, M. E. ; Vuori, K. ; Milde, T. ; Cho, Y.-J. ; Li, X.-N. ; Olson, J. M. ; Reyes, I. ; Snuderl, M. ; Wong, T. C. ; Dimmock, D. P. ; Nahas, S. A. ; Malicki, D. ; Crawford, J. R. ; Levy, M. L. ; Van Allen, E. M. ; Pfister, S. ; Tamayo, P. ; Kool, M. ; Mesirov, J. P. ; Wechsler-Reya, R. J.
Selt, F. ; van Tilburg, C. M. ; Bison, B. ; Sievers, P. ; Harting, I. ; Ecker, J. ; Pajtler, K. W. ; Sahm, F. ; Bahr, A. ; Simon, M. ; Jones, D. T. W. ; Well, L. ; Mautner, V.-F. ; Capper, D. ; Hernáiz Driever, P. ; Gnekow, A. ; Pfister, S. M. ; Witt, O. ; Milde, T.
Fittall, M. W. ; Lyskjaer, I. ; Ellery, P. ; Lombard, P. ; Ijaz, J. ; Strobl, A.-C. ; Oukrif, D. ; Tarabichi, M. ; Sill, M. ; Koelsche, C. ; Mechtersheimer, G. ; Demeulemeester, J. ; Tirabosco, R. ; Amary, F. ; Campbell, P. J. ; Pfister, S. ; Jones, D. T. W. ; Pillay, N. ; Van Loo, P. ; Behjati, S. ; Flanagan, A. M.
Kutscher, L. ; Okonechnikov, K. ; Batora, N. ; Clark, J. ; Silva, P. B. G. ; Vouri, M. ; van Rijn, S. ; Sieber, L. ; Statz, B. ; Gearhart, M. D. ; Shiraishi, R. ; Mack, N. ; Orr, B. A. ; Korshunov, A. ; Gudenas, B. L. ; Smith, K. S. ; Mercier, A. L. ; Ayrault, O. ; Hoshino, M. ; Kool, M. ; von Hoff, K. ; Graf, N. ; Fleishhack, G. ; Bardwell, V. J. ; Pfister, S. M. ; Northcott, P. A. ; Kawauchi, D.
Banan, R. ; Stichel, D. ; Bleck, A. ; Hong, B. ; Lehmann, U. ; Suwala, A. ; Reinhardt, A. ; Schrimpf, D. ; Buslei, R. ; Stadelmann, C. ; Ehlert, K. ; Prinz, M. ; Acker, T. ; Schittenhelm, J. ; Kaul, D. ; Schweizer, L. ; Capper, D. ; Harter, P. ; Etminan, N. ; Jones, D. T. W. ; Pfister, S. M. ; Herold-Mende, C. ; Wick, W. ; Sahm, F. ; von Deimling, A. ; Hartmann, C. ; Reuß, D.
Susanto, E. ; Marin Navarro, A. ; Zhou, L. ; Sundström, A. ; van Bree, N. ; Stantic, M. ; Moslem, M. ; Tailor, J. ; Rietdijk, J. ; Zubillaga, V. ; Hübner, J.-M. ; Weishaupt, H. ; Wolfsberger, J. ; Alafuzoff, I. ; Nordgren, A. ; Magnaldo, T. ; Siesjö, P. ; Johnsen, J. I. ; Kool, M. ; Tammimies, K. ; Darabi, A. ; Swartling, F. J. ; Falk, A. ; Wilhelm, M.
Sánchez-Maldonado, J. M. ; Campa, D. ; Springer, J. ; Badiola, J. ; Niazi, Y. ; Moñiz-Díez, A. ; Hernández-Mohedo, F. ; González-Sierra, P. ; Ter Horst, R. ; Macauda, A. ; Brezina, S. ; Cunha, C. ; Lackner, M. ; López-Nevot, M. A. ; Fianchi, L. ; Pagano, L. ; López-Fernández, E. ; Potenza, L. ; Luppi, M. ; Moratalla, L. ; Rodríguez-Sevilla, J. J. ; Fonseca, J. E. ; Tormo, M. ; Solano, C. ; Clavero, E. ; Romero, A. ; Li, Y. ; Lass-Flörl, C. ; Einsele, H. ; Vazquez, L. ; Loeffler, J. ; Hemminki, K. ; Carvalho, A. ; Netea, M. G. ; Gsur, A. ; Dumontet, C. ; Canzian, F. ; Försti, A. ; Jurado, M. ; Sainz, J.
Gojo, J. ; Englinger, B. ; Jiang, L. ; Hübner, J. M. ; Shaw, M. L. ; Hack, O. A. ; Madlener, S. ; Kirchhofer, D. ; Liu, I. ; Pyrdol, J. ; Hovestadt, V. ; Mazzola, E. ; Mathewson, N. D. ; Trissal, M. ; Lötsch, D. ; Dorfer, C. ; Haberler, C. ; Halfmann, A. ; Mayr, L. ; Peyrl, A. ; Geyeregger, R. ; Schwalm, B. ; Mauermann, M. ; Pajtler, K. W. ; Milde, T. ; Shore, M. E. ; Geduldig, J. E. ; Pelton, K. ; Czech, T. ; Ashenberg, O. ; Wucherpfennig, K. W. ; Rozenblatt-Rosen, O. ; Alexandrescu, S. ; Ligon, K. L. ; Pfister, S. M. ; Regev, A. ; Slavc, I. ; Berger, W. ; Suvà, M. L. ; Kool, M. ; Filbin, M. G.
Sommerkamp, A. ; Uhrig, S. ; Stichel, D. ; St-Onge, P. ; Sun, P. ; Jäger, N. ; von Deimling, A. ; Sahm, F. ; Pfister, S. M. ; Korshunov, A. ; Sinnett, D. ; Jabado, N. ; Wefers, A. K. ; Jones, D.
Sievers, P. ; Hielscher, T. ; Schrimpf, D. ; Stichel, D. ; Reuss, D. E. ; Berghoff, A. S. ; Neidert, M. C. ; Wirsching, H.-G. ; Mawrin, C. ; Ketter, R. ; Paulus, W. ; Reifenberger, G. ; Lamszus, K. ; Westphal, M. ; Etminan, N. ; Ratliff, M. ; Herold-Mende, C. ; Pfister, S. M. ; Jones, D. T. W. ; Weller, M. ; Harter, P. N. ; Wick, W. ; Preusser, M. ; von Deimling, A. ; Sahm, F.
Usta, D. ; Sigaud, R. ; Buhl, J. L. ; Selt, F. ; Marquardt, V. ; Pauck, D. ; Jansen, J. ; Pusch, S. ; Ecker, J. ; Hielscher, T. ; Vollmer, J. ; Sommerkamp, A. C. ; Rubner, T. ; Hargrave, D. ; van Tilburg, C. M. ; Pfister, S. M. ; Jones, D. T. W. ; Remke, M. ; Brummer, T. ; Witt, O. ; Milde, T.
Weber, J. ; Braun, C. J. ; Saur, D. ; Rad, R.
van Tilburg, C. M. ; Witt, R. ; Heiss, M. ; Pajtler, K. W. ; Plass, C. ; Poschke, I. ; Platten, M. ; Harting, I. ; Sedlaczek, O. ; Freitag, A. ; Meyrath, D. ; Taylor, L. ; Balasubramanian, G. P. ; Jäger, N. ; Pfaff, E. ; Jones, B. C. ; Milde, T. ; Pfister, S. M. ; Jones, D. T. W. ; Kopp-Schneider, A. ; Witt, O.
Theruvath, J. ; Sotillo, E. ; Mount, C. W. ; Graef, C. M. ; Delaidelli, A. ; Heitzeneder, S. ; Labanieh, L. ; Dhingra, S. ; Leruste, A. ; Majzner, R. G. ; Xu, P. ; Mueller, S. ; Yecies, D. W. ; Finetti, M. A. ; Williamson, D. ; Johann, P. D. ; Kool, M. ; Pfister, S. ; Hasselblatt, M. ; Frühwald, M. C. ; Delattre, O. ; Surdez, D. ; Bourdeaut, F. ; Puget, S. ; Zaidi, S. ; Mitra, S. S. ; Cheshier, S. ; Sorensen, P. H. ; Monje, M. ; Mackall, C. L.
Mynarek, M. ; von Hoff, K. ; Pietsch, T. ; Ottensmeier, H. ; Warmuth-Metz, M. ; Bison, B. ; Pfister, S. ; Korshunov, A. ; Sharma, T. ; Jaeger, N. ; Ryzhova, M. ; Zheludkova, O. ; Golanov, A. ; Rushing, E. J. ; Hasselblatt, M. ; Koch, A. ; Schüller, U. ; von Deimling, A. ; Sahm, F. ; Sill, M. ; Riemenschneider, M. J. ; Dohmen, H. ; Monoranu, C. M. ; Sommer, C. ; Staszewski, O. ; Mawrin, C. ; Schittenhelm, J. ; Brück, W. ; Filipski, K. ; Hartmann, C. ; Meinhardt, M. ; Pietschmann, K. ; Haberler, C. ; Slavc, I. ; Gerber, N. U. ; Grotzer, M. ; Benesch, M. ; Schlegel, P. G. ; Deinlein, F. ; von Bueren, A. O. ; Friedrich, C. ; Juhnke, B.-O. ; Obrecht, D. ; Fleischhack, G. ; Kwiecien, R. ; Faldum, A. ; Kortmann, R. D. ; Kool, M. ; Rutkowski, S.
Schubert, N. A. ; Lowery, C. D. ; Bergthold, G. ; Koster, J. ; Eleveld, T. F. ; Rodríguez, A. ; Jones, D. T. W. ; German Cancer Research Center , H. ; Vassal, G. ; Stancato, L. F. ; Pfister, S. M. ; German Cancer Research Center , H. ; Heidelberg University Hospital, H. ; Caron, H. N. ; Molenaar, J. J.
Calandrini, C. ; Schutgens, F. ; Oka, R. ; Margaritis, T. ; Candelli, T. ; Mathijsen, L. ; Ammerlaan, C. ; van Ineveld, R. L. ; Derakhshan, S. ; de Haan, S. ; Dolman, E. ; Lijnzaad, P. ; Custers, L. ; Begthel, H. ; Kerstens, H. H. D. ; Visser, L. L. ; Rookmaaker, M. ; Verhaar, M. ; Tytgat, G. A. M. ; Kemmeren, P. ; de Krijger, R. R. ; Al-Saadi, R. ; Pritchard-Jones, K. ; Kool, M. ; Rios, A. C. ; van den Heuvel-Eibrink, M. M. ; Molenaar, J. J. ; van Boxtel, R. ; Holstege, F. C. P. ; Clevers, H. ; Drost, J.
Horak, P. ; Uhrig, S. ; Witzel, M. ; Gil-Farina, I. ; Hutter, B. ; Rath, T. ; Gieldon, L. ; Balasubramanian, G. P. ; Pastor, X. ; Heilig, C. E. ; Richter, D. ; Schröck, E. ; Ball, C. R. ; Brors, B. ; Braun, C. J. ; Albert, M. H. ; Scholl, C. ; von Kalle, C. ; Schmidt, M. ; Fröhling, S. ; Klein, C. ; Glimm, H.
Hong, D. S. ; DuBois, S. G. ; Kummar, S. ; Farago, A. F. ; Albert, C. M. ; Rohrberg, K. S. ; van Tilburg, C. M. ; Nagasubramanian, R. ; Berlin, J. D. ; Federman, N. ; Mascarenhas, L. ; Geoerger, B. ; Dowlati, A. ; Pappo, A. S. ; Bielack, S. ; Doz, F. ; McDermott, R. ; Patel, J. D. ; Schilder, R. J. ; Tahara, M. ; Pfister, S. M. ; Witt, O. ; Ladanyi, M. ; Rudzinski, E. R. ; Nanda, S. ; Childs, B. H. ; Laetsch, T. W. ; Hyman, D. M. ; Drilon, A.
Gerstung, M. ; Jolly, C. ; Leshchiner, I. ; Dentro, S. ; Gonzalez, S. ; Rosebrock, D. ; Mitchell, T. J. ; Rubanova, Y. ; Anur, P. ; Yu, K. ; Tarabichi, M. ; Deshwar, A. ; Wintersinger, J. ; Kleinheinz, K. ; Vázquez-García, I. ; Haase, K. ; Jerman, L. ; Sengupta, S. ; Macintyre, G. ; Malikic, S. ; Donmez, N. ; Livitz, D. G. ; Cmero, M. ; Demeulemeester, J. ; Schumacher, S. ; Fan, Y. ; Yao, X. ; Lee, J. ; Schlesner, M. ; Boutros, P. C. ; Bowtell, D. D. ; Zhu, H. ; Getz, G. ; Imielinski, M. ; Beroukhim, R. ; Sahinalp, S. C. ; Ji, Y. ; Peifer, M. ; Markowetz, F. ; Mustonen, V. ; Yuan, K. ; Wang, W. ; Morris, Q. D. ; PCAWGEvolution&HeterogeneityWorkingGroup ; Spellman, P. T. ; Wedge, D. C. ; Van Loo, P. ; PCAWGConsortium
Frühwald, M. C. ; Hasselblatt, M. ; Nemes, K. ; Bens, S. ; Steinbügl, M. ; Johann, P. D. ; Kerl, K. ; Hauser, P. ; Quiroga, E. ; Solano-Paez, P. ; Biassoni, V. ; Gil-da-Costa, M. J. ; Perek-Polnik, M. ; van de Wetering, M. ; Sumerauer, D. ; Pears, J. ; Stabell, N. ; Holm, S. ; Hengartner, H. ; Gerber, N. U. ; Grotzer, M. ; Boos, J. ; Ebinger, M. ; Tippelt, S. ; Paulus, W. ; Furtwängler, R. ; Hernáiz-Driever, P. ; Reinhard, H. ; Rutkowski, S. ; Schlegel, P.-G. ; Schmid, I. ; Kortmann, R.-D. ; Timmermann, B. ; Warmuth-Metz, M. ; Kordes, U. ; Gerss, J. ; Nysom, K. ; Schneppenheim, R. ; Siebert, R. ; Kool, M. ; Graf, N.
Deng, M. Y. ; Sill, M. ; Sturm, D. ; Stichel, D. ; Witt, H. ; Ecker, J. ; Wittmann, A. ; Schittenhelm, J. ; Ebinger, M. ; Schuhmann, M. U. ; Figarella-Branger, D. ; Aronica, E. ; Staszewski, O. ; Preusser, M. ; Haberler, C. ; Lauten, M. ; Schüller, U. ; Hartmann, C. ; Snuderl, M. ; Dunham, C. ; Jabado, N. ; Wesseling, P. ; Deckert, M. ; Keyvani, K. ; Gottardo, N. ; Giangaspero, F. ; von Hoff, K. ; Ellison, D. W. ; Pietsch, T. ; Herold Mende, C. ; Milde, T. ; Witt, O. ; Kool, M. ; Korshunov, A. ; Wick, W. ; von Deimling, A. ; Pfister, S. M. ; Jones, D. T. W. ; Sahm, F.
Kommoss, F. K. F. ; Stichel, D. ; Schrimpf, D. ; Kriegsmann, M. ; Tessier-Cloutier, B. ; Talhouk, A. ; McAlpine, J. N. ; Chang, K. T. E. ; Sturm, D. ; Pfister, S. M. ; Romero-Pérez, L. ; Kirchner, T. ; Grünewald, T. G. P. ; Buslei, R. ; Sinn, H.-P. ; Mechtersheimer, G. ; Schirmacher, P. ; Schmidt, D. ; Lehr, H.-A. ; Sahm, F. ; Huntsman, D. G. ; Gilks, C. B. ; Kommoss, F. ; von Deimling, A. ; Koelsche, C.
Pickles, J. C. ; Fairchild, A. R. ; Stone, T. J. ; Brownlee, L. ; Merve, A. ; Yasin, S. A. ; Avery, A. ; Ahmed, S. W. ; Ogunbiyi, O. ; Gonzalez Zapata, J. ; Peary, A. F. ; Edwards, M. ; Wilkhu, L. ; Dryden, C. ; Ladon, D. ; Kristiansen, M. ; Rowe, C. ; Kurian, K. M. ; Nicoll, J. A. R. ; Mitchell, C. ; Bloom, T. ; Hilton, D. A. ; Al-Sarraj, S. ; Doey, L. ; Johns, P. N. ; Bridges, L. R. ; Chakrabarty, A. ; Ismail, A. ; Rathi, N. ; Syed, K. ; Lammie, G. A. ; Limback-Stanic, C. ; Smith, C. ; Torgersen, A. ; Rae, F. ; Hill, R. M. ; Clifford, S. C. ; Grabovska, Y. ; Williamson, D. ; Clarke, M. ; Jones, C. ; Capper, D. ; Sill, M. ; von Deimling, A. ; Pfister, S. M. ; Jones, D. T. W. ; Hargrave, D. ; Chalker, J. ; Jacques, T. S.
Korshunov, A. ; Okonechnikov, K. ; Sahm, F. ; Ryzhova, M. ; Stichel, D. ; Schrimpf, D. ; Ghasemi, D. R. ; Pajtler, K. W. ; Antonelli, M. ; Donofrio, V. ; Mastronuzzi, A. ; Rossi, S. ; Camassei, F. D. ; Buccoliero, A. M. ; Haberler, C. ; Slavc, I. ; Dahiya, S. ; Casalini, B. ; Sievers, P. ; Meyer, J. ; Kumirova, E. ; Zheludkova, O. ; Golanov, A. ; Jones, D. T. W. ; Pfister, S. M. ; Kool, M. ; von Deimling, A.
Pfaff, E. ; Aichmüller, C. ; Sill, M. ; Stichel, D. ; Snuderl, M. ; Karajannis, M. A. ; Schuhmann, M. U. ; Schittenhelm, J. ; Hasselblatt, M. ; Thomas, C. ; Korshunov, A. ; Rhizova, M. ; Wittmann, A. ; Kaufhold, A. ; Iskar, M. ; Ketteler, P. ; Lohmann, D. ; Orr, B. A. ; Ellison, D. W. ; von Hoff, K. ; Mynarek, M. ; Rutkowski, S. ; Sahm, F. ; von Deimling, A. ; Lichter, P. ; Kool, M. ; Zapatka, M. ; Pfister, S. M. ; Jones, D.
Bleeke, M. ; Johann, P. ; Gröbner, S. ; Alten, J. ; Cario, G. ; Schäfer, H. ; Klapper, W. ; Khoury, J. ; Pfister, S. ; Müller, I.
Sievers, P. ; Chiang, J. ; Schrimpf, D. ; Stichel, D. ; Paramasivam, N. ; Sill, M. ; Gayden, T. ; Casalini, B. ; Reuss, D. E. ; Dalton, J. ; Pajtler, K. W. ; Hänggi, D. ; Herold-Mende, C. ; Rushing, E. ; Korshunov, A. ; Mawrin, C. ; Weller, M. ; Schlesner, M. ; Wick, W. ; Jabado, N. ; Jones, D. T. W. ; Pfister, S. M. ; von Deimling, A. ; Ellison, D. W. ; Sahm, F.
Thomas, C. ; Wefers, A. ; Bens, S. ; Nemes, K. ; Agaimy, A. ; Oyen, F. ; Vogelgesang, S. ; Rodriguez, F. J. ; Brett, F. M. ; McLendon, R. ; Bodi, I. ; Burel-Vandenbos, F. ; Keyvani, K. ; Tippelt, S. ; Poulsen, F. R. ; Lipp, E. S. ; Giannini, C. ; Reifenberger, G. ; Kuchelmeister, K. ; Pietsch, T. ; Kordes, U. ; Siebert, R. ; Frühwald, M. C. ; Johann, P. D. ; Sill, M. ; Kool, M. ; von Deimling, A. ; Paulus, W. ; Hasselblatt, M.
Chattopadhyay, S. ; Thomsen, H. ; Weinhold, N. ; Meziane, I. ; Huhn, S. ; da Silva Filho, M. I. ; Vodicka, P. ; Vodickova, L. ; Hoffmann, P. ; Nöthen, M. M. ; Jöckel, K.-H. ; Schmidt, B. ; Landi, S. ; Hajek, R. ; Hallmans, G. ; Pettersson-Kymmer, U. ; Ohlsson, C. ; Milani, P. ; Merlini, G. ; Rowcieno, D. ; Hawkins, P. ; Hegenbart, U. ; Palladini, G. ; Wechalekar, A. ; Schönland, S. O. ; Houlston, R. ; Goldschmidt, H. ; Hemminki, K. ; Försti, A.
Begemann, M. ; Waszak, S. M. ; Robinson, G. W. ; Jäger, N. ; Sharma, T. ; Knopp, C. ; Kraft, F. ; Moser, O. ; Mynarek, M. ; Guerrini-Rousseau, L. ; Brugieres, L. ; Varlet, P. ; Pietsch, T. ; Bowers, D. C. ; Chintagumpala, M. ; Sahm, F. ; Korbel, J. O. ; Rutkowski, S. ; Eggermann, T. ; Gajjar, A. ; Northcott, P. ; Elbracht, M. ; Pfister, S. M. ; Kontny, U. ; Kurth, I.
Benesch, M. ; Nemes, K. ; Neumayer, P. ; Hasselblatt, M. ; Timmermann, B. ; Bison, B. ; Ebetsberger-Dachs, G. ; Bourdeaut, F. ; Dufour, C. ; Biassoni, V. ; Morales La Madrid, A. ; Entz-Werle, N. ; Laithier, V. ; Quehenberger, F. ; Weis, S. ; Sumerauer, D. ; Siebert, R. ; Bens, S. ; Schneppenheim, R. ; Kool, M. ; Modena, P. ; Fouyssac, F. ; C Frühwald, M.
Jabarkheel, R. ; Amayiri, N. ; Yecies, D. ; Huang, Y. ; Toescu, S. ; Nobre, L. ; Mabbott, D. J. ; Sudhakar, S. V. ; Malik, P. ; Laughlin, S. ; Swaidan, M. ; Al Hussaini, M. ; Musharbash, A. ; Chacko, G. ; Mathew, L. G. ; Fisher, P. G. ; Hargrave, D. ; Bartels, U. ; Tabori, U. ; Pfister, S. M. ; Aquilina, K. ; Taylor, M. D. ; Grant, G. A. ; Bouffet, E. ; Mankad, K. ; Yeom, K. W. ; Ramaswamy, V.
Reisinger, D. ; Gojo, J. ; Kasprian, G. ; Haberler, C. ; Peyrl, A. ; Azizi, A. A. ; Mayr, L. ; Chocholous, M. ; Kool, M. ; Czech, T. ; Slavc, I.
van Tilburg, C. M. ; Milde, T. ; Witt, R. ; Ecker, J. ; Hielscher, T. ; Seitz, A. ; Schenk, J.-P. ; Buhl, J. L. ; Riehl, D. ; Frühwald, M. C. ; Pekrun, A. ; Rossig, C. ; Wieland, R. ; Flotho, C. ; Kordes, U. ; Gruhn, B. ; Simon, T. ; Linderkamp, C. ; Sahm, F. ; Taylor, L. ; Freitag, A. ; Burhenne, J. ; Foerster, K. I. ; Meid, A. D. ; Pfister, S. M. ; Karapanagiotou-Schenkel, I. ; Witt, O.
Franceschi, E. ; Hofer, S. ; Brandes, A. A. ; Frappaz, D. ; Kortmann, R.-D. ; Bromberg, J. ; Dangouloff-Ros, V. ; Boddaert, N. ; Hattingen, E. ; Wiestler, B. ; Clifford, S. C. ; Figarella-Branger, D. ; Giangaspero, F. ; Haberler, C. ; Pietsch, T. ; Pajtler, K. W. ; Pfister, S. M. ; Guzman, R. ; Stummer, W. ; Combs, S. E. ; Seidel, C. ; Beier, D. ; McCabe, M. G. ; Grotzer, M. ; Laigle-Donadey, F. ; Stücklin, A. S. G. ; Idbaih, A. ; Preusser, M. ; van den Bent, M. ; Weller, M. ; Hau, P.
Aldape, K. ; Brindle, K. M. ; Chesler, L. ; Chopra, R. ; Gajjar, A. ; Gilbert, M. R. ; Gottardo, N. ; Gutmann, D. H. ; Hargrave, D. ; Holland, E. C. ; Jones, D. T. W. ; Joyce, J. A. ; Kearns, P. ; Kieran, M. W. ; Mellinghoff, I. K. ; Merchant, M. ; Pfister, S. M. ; Pollard, S. M. ; Ramaswamy, V. ; Rich, J. N. ; Robinson, G. W. ; Rowitch, D. H. ; Sampson, J. H. ; Taylor, M. D. ; Workman, P. ; Gilbertson, R. J.
Aldape, K. ; Brindle, K. M. ; Chesler, L. ; Chopra, R. ; Gajjar, A. ; Gilbert, M. R. ; Gottardo, N. ; Gutmann, D. H. ; Hargrave, D. ; Holland, E. C. ; Jones, D. T. W. ; Joyce, J. A. ; Kearns, P. ; Kieran, M. W. ; Mellinghoff, I. K. ; Merchant, M. ; Pfister, S. M. ; Pollard, S. M. ; Ramaswamy, V. ; Rich, J. N. ; Robinson, G. W. ; Rowitch, D. H. ; Sampson, J. H. ; Taylor, M. D. ; Workman, P. ; Gilbertson, R. J.
Treisman, D. M. ; Li, Y. ; Pierce, B. R. ; Li, C. ; Chervenak, A. P. ; Tomasek, G. J. ; Lozano, G. ; Zheng, X. ; Kool, M. ; Zhu, Y.
Chun, H. E. ; Johann, P. D. ; Milne, K. ; Zapatka, M. ; Buellesbach, A. ; Ishaque, N. ; Iskar, M. ; Erkek, S. ; Wei, L. ; Tessier-Cloutier, B. ; Lever, J. ; Titmuss, E. ; Topham, J. T. ; Bowlby, R. ; Chuah, E. ; Mungall, K. L. ; Ma, Y. ; Mungall, A. J. ; Moore, R. A. ; Taylor, M. D. ; Gerhard, D. S. ; Jones, S. J. M. ; Korshunov, A. ; Gessler, M. ; Kerl, K. ; Hasselblatt, M. ; Frühwald, M. C. ; Perlman, E. J. ; Nelson, B. H. ; Pfister, S. M. ; Marra, M. A. ; Kool, M.
Jaju, A. ; Hwang, E. I. ; Kool, M. ; Capper, D. ; Chavez, L. ; Brabetz, S. ; Billups, C. ; Li, Y. ; Fouladi, M. ; Packer, R. J. ; Pfister, S. M. ; Olson, J. M. ; Heier, L. A.
Senfter, D. ; Samadaei, M. ; Mader, R. M. ; Gojo, J. ; Peyrl, A. ; Krupitza, G. ; Kool, M. ; Sill, M. ; Haberler, C. ; Ricken, G. ; Czech, T. ; Slavc, I. ; Madlener, S.
Pajtler, K. ; Wei, Y. ; Okonechnikov, K. ; Silva, P. B. G. ; Vouri, M. ; Zhang, L. ; Brabetz, S. ; Sieber, L. ; Gulley, M. ; Mauermann, M. ; Wedig, T. ; Mack, N. ; Imamura Kawasawa, Y. ; Sharma, T. ; Zuckermann, M. ; Andreiuolo, F. ; Holland, E. ; Maass, K. ; Körkel-Qu, H. ; Liu, H.-K. ; Sahm, F. ; Capper, D. ; Bunt, J. ; Richards, L. J. ; Jones, D. T. W. ; Korshunov, A. ; Chavez, L. ; Lichter, P. ; Hoshino, M. ; Pfister, S. ; Kool, M. ; Li, W. ; Kawauchi, D.
Pienkowska, M. ; Choufani, S. ; Turinsky, A. L. ; Guha, T. ; Merino, D. M. ; Novokmet, A. ; Brudno, M. ; Weksberg, R. ; Shlien, A. ; Hawkins, C. ; Bouffet, E. ; Tabori, U. ; Gilbertson, R. ; Finlay, J. L. ; Jabado, N. ; Thomas, C. ; Sill, M. ; Capper, D. ; Hasselblatt, M. ; Malkin, D.
Ghasemi, D. R. ; Sill, M. ; Okonechnikov, K. ; Korshunov, A. ; Yip, S. ; Schutz, P. W. ; Scheie, D. ; Kruse, A. ; Harter, P. N. ; Kastelan, M. ; Wagner, M. ; Hartmann, C. ; Benzel, J. ; Maass, K. K. ; Khasraw, M. ; Sträter, R. ; Thomas, C. ; Paulus, W. ; Kratz, C. P. ; Witt, H. ; Kawauchi, D. ; Herold-Mende, C. ; Sahm, F. ; Brandner, S. ; Kool, M. ; Jones, D. T. W. ; von Deimling, A. ; Pfister, S. M. ; Reuss, D. E. ; Pajtler, K.
Stichel, D. ; Schrimpf, D. ; Casalini, B. ; Meyer, J. ; Wefers, A. K. ; Sievers, P. ; Korshunov, A. ; Koelsche, C. ; Reuss, D. E. ; Reinhardt, A. ; Ebrahimi, A. ; Fernández-Klett, F. ; Kessler, T. ; Sturm, D. ; Ecker, J. ; Milde, T. ; Herold-Mende, C. ; Witt, O. ; Pfister, S. M. ; Wick, W. ; Jones, D. T. W. ; von Deimling, A. ; Sahm, F.
Sievers, P. ; Appay, R. ; Schrimpf, D. ; Stichel, D. ; Reuss, D. E. ; Wefers, A. K. ; Reinhardt, A. ; Coras, R. ; Ruf, V. C. ; Schmid, S. ; de Stricker, K. ; Boldt, H. B. ; Kristensen, B. W. ; Petersen, J. K. ; Ulhøi, B. P. ; Gardberg, M. ; Aronica, E. ; Hasselblatt, M. ; Brück, W. ; Bielle, F. ; Mokhtari, K. ; Lhermitte, B. ; Wick, W. ; Herold-Mende, C. ; Hänggi, D. ; Brandner, S. ; Giangaspero, F. ; Capper, D. ; Rushing, E. ; Wesseling, P. ; Pfister, S. M. ; Figarella-Branger, D. ; von Deimling, A. ; Sahm, F. ; Jones, D.
Park, A. K. ; Lee, J. Y. ; Cheong, H. ; Ramaswamy, V. ; Park, S.-H. ; Kool, M. ; Phi, J. H. ; Choi, S. A. ; Cavalli, F. ; Taylor, M. D. ; Kim, S.-K.
Huang, M. ; Tailor, J. ; Zhen, Q. ; Gillmor, A. H. ; Miller, M. L. ; Weishaupt, H. ; Chen, J. ; Zheng, T. ; Nash, E. K. ; McHenry, L. K. ; An, Z. ; Ye, F. ; Takashima, Y. ; Clarke, J. ; Ayetey, H. ; Cavalli, F. M. G. ; Luu, B. ; Moriarity, B. S. ; Ilkhanizadeh, S. ; Chavez, L. ; Yu, C. ; Kurian, K. M. ; Magnaldo, T. ; Sevenet, N. ; Koch, P. ; Pollard, S. M. ; Dirks, P. ; Snyder, M. P. ; Largaespada, D. A. ; Cho, Y. J. ; Phillips, J. J. ; Swartling, F. J. ; Morrissy, A. S. ; Kool, M. ; Pfister, S. M. ; Taylor, M. D. ; Smith, A. ; Weiss, W. A.
Fangusaro, J. ; Onar-Thomas, A. ; Young Poussaint, T. ; Wu, S. ; Ligon, A. H. ; Lindeman, N. ; Banerjee, A. ; Packer, R. J. ; Kilburn, L. B. ; Goldman, S. ; Pollack, I. F. ; Qaddoumi, I. ; Jakacki, R. I. ; Fisher, P. G. ; Dhall, G. ; Baxter, P. ; Kreissman, S. G. ; Stewart, C. F. ; Jones, D. T. W. ; Pfister, S. M. ; Vezina, G. ; Stern, J. S. ; Panigrahy, A. ; Patay, Z. ; Tamrazi, B. ; Jones, J. Y. ; Haque, S. S. ; Enterline, D. S. ; Cha, S. ; Fisher, M. J. ; Doyle, L. A. ; Smith, M. ; Dunkel, I. J. ; Fouladi, M.
Schramm, K. ; Iskar, M. ; Statz, B. ; Jäger, N. ; Haag, D. ; Słabicki, M. ; Pfister, S. M. ; Zapatka, M. ; Gronych, J. ; Jones, D. T. W. ; Lichter, P.
Pfaff, E. ; El Damaty, A. ; Balasubramanian, G. P. ; Blattner-Johnson, M. ; Worst, B. C. ; Stark, S. ; Witt, H. ; Pajtler, K. W. ; van Tilburg, C. M. ; Witt, R. ; Milde, T. ; Jakobs, M. ; Fiesel, P. ; Frühwald, M. C. ; Hernáiz Driever, P. ; Thomale, U. W. ; Schuhmann, M. U. ; Metzler, M. ; Bochennek, K. ; Simon, T. ; Dürken, M. ; Karremann, M. ; Knirsch, S. ; Ebinger, M. ; von Bueren, A. O. ; Pietsch, T. ; Herold-Mende, C. ; Reuss, D. E. ; Kiening, K. ; Lichter, P. ; Eggert, A. ; Kramm, C. M. ; Pfister, S. M. ; Jones, D. T. W. ; Bächli, H. ; Witt, O.
Binder, H. ; Willscher, E. ; Loeffler-Wirth, H. ; Hopp, L. ; Jones, D. T. W. ; Pfister, S. M. ; Kreuz, M. ; Gramatzki, D. ; Fortenbacher, E. ; Hentschel, B. ; Tatagiba, M. ; Herrlinger, U. ; Vatter, H. ; Matschke, J. ; Westphal, M. ; Krex, D. ; Schackert, G. ; Tonn, J. C. ; Schlegel, U. ; Steiger, H.-J. ; Wick, W. ; Weber, R. G. ; Weller, M. ; Loeffler, M.
Wirthschaft, P. ; Bode, J. ; Soni, H. ; Dietrich, F. ; Krüwel, T. ; Fischer, B. ; Knobbe-Thomsen, C. B. ; Rossetti, G. ; Hentschel, A. ; Mack, N. ; Schönig, K. ; Breckwoldt, M. O. ; Schmandke, A. ; Pusch, S. ; Medenbach, J. ; Bendszus, M. ; Schwab, M. E. ; von Deimling, A. ; Kool, M. ; Herold-Mende, C. ; Reifenberger, G. ; Ahrends, R. ; Tews, B.
Paramasivam, N. ; Hübschmann, D. ; Toprak, U. H. ; Ishaque, N. ; Neidert, M. ; Schrimpf, D. ; Stichel, D. ; Reuss, D. ; Sievers, P. ; Reinhardt, A. ; Wefers, A. K. ; Jones, D. T. W. ; Gu, Z. ; Werner, J. ; Uhrig, S. ; Wirsching, H.-G. ; Schick, M. ; Bewerunge-Hudler, M. ; Beck, K. ; Brehmer, S. ; Urbschat, S. ; Seiz-Rosenhagen, M. ; Hänggi, D. ; Herold-Mende, C. ; Ketter, R. ; Eils, R. ; Ram, Z. ; Pfister, S. M. ; Wick, W. ; Weller, M. ; Grossmann, R. ; von Deimling, A. ; Schlesner, M. ; Sahm, F.
Thomas, C. ; Knerlich-Lukoschus, F. ; Reinhard, H. ; Johann, P. D. ; Sturm, D. ; Sahm, F. ; Bens, S. ; Vogt, J. ; Nemes, K. ; Oyen, F. ; Kordes, U. ; Siebert, R. ; Schneppenheim, R. ; Messing-Jünger, M. ; Pietsch, T. ; von Deimling, A. ; Paulus, W. ; Pfister, S. M. ; Kool, M. ; Frühwald, M. C. ; Hasselblatt, M.
Hou, Y. ; Pinheiro, J. ; Sahm, F. ; Reuss, D. E. ; Schrimpf, D. ; Stichel, D. ; Casalini, B. ; Koelsche, C. ; Sievers, P. ; Wefers, A. K. ; Reinhardt, A. ; Ebrahimi, A. ; Fernández-Klett, F. ; Pusch, S. ; Meier, J. ; Schweizer, L. ; Paulus, W. ; Prinz, M. ; Hartmann, C. ; Plate, K. H. ; Reifenberger, G. ; Pietsch, T. ; Varlet, P. ; Pagès, M. ; Schüller, U. ; Scheie, D. ; de Stricker, K. ; Frank, S. ; Hench, J. ; Pollo, B. ; Brandner, S. ; Unterberg, A. ; Pfister, S. M. ; Jones, D. T. W. ; Korshunov, A. ; Wick, W. ; Capper, D. ; Blümcke, I. ; von Deimling, A. ; Bertero, L.
Sharma, T. ; Schwalbe, E. C. ; Williamson, D. ; Sill, M. ; Hovestadt, V. ; Mynarek, M. ; Rutkowski, S. ; Robinson, G. W. ; Gajjar, A. ; Cavalli, F. ; Ramaswamy, V. ; Taylor, M. D. ; Lindsey, J. C. ; Hill, R. M. ; Jäger, N. ; Korshunov, A. ; Hicks, D. ; Bailey, S. ; Kool, M. ; Chavez, L. ; Northcott, P. A. ; Pfister, S. ; Clifford, S. C.
Pagès, M. ; Pajtler, K. ; Puget, S. ; Castel, D. ; Boddaert, N. ; Tauziède‐Espariat, A. ; Picot, S. ; Debily, M. ; Kool, M. ; Capper, D. ; Sainte‐Rose, C. ; Chrétien, F. ; Pfister, S. ; Pietsch, T. ; Grill, J. ; Varlet, P. ; Andreiuolo, F.
Benesch, M. ; Mynarek, M. ; Witt, H. ; Warmuth-Metz, M. ; Pietsch, T. ; Bison, B. ; Pfister, S. ; Pajtler, K. ; Kool, M. ; Schüller, U. ; Pietschmann, K. ; Juhnke, B.-O. ; Tippelt, S. ; Fleischhack, G. ; Schmid, I. ; Kramm, C. M. ; Vorwerk, P. ; Beilken, A. ; Classen, C. F. ; Hernáiz Driever, P. ; Kropshofer, G. ; Imschweiler, T. ; Lemmer, A. ; Kortmann, R.-D. ; Rutkowski, S. ; von Hoff, K.
Tao, R. ; Murad, N. ; Xu, Z. ; Zhang, P. ; Okonechnikov, K. ; Kool, M. ; Rivero-Hinojosa, S. ; Lazarski, C. ; Zheng, P. ; Liu, Y. ; Eberhart, C. G. ; Rood, B. R. ; Packer, R. ; Pei, Y.
Koelsche, C. ; Kriegsmann, M. ; Kommoss, F. K. F. ; Stichel, D. ; Kriegsmann, K. ; Vokuhl, C. ; Grünewald, T. G. P. ; Romero-Pérez, L. ; Kirchner, T. ; de Alava, E. ; Diaz-Martin, J. ; Hartmann, W. ; Baumhoer, D. ; Antonescu, C. R. ; Szuhai, K. ; Flucke, U. ; Dirksen, U. ; Pfister, S. ; Jones, D. ; Mechtersheimer, G. ; von Deimling, A.
Körber, V. ; Yang, J. ; Barah, P. ; Wu, Y. ; Stichel, D. ; Gu, Z. ; Fletcher, M. ; Jones, D. ; Hentschel, B. ; Lamszus, K. ; Tonn, J. C. ; Schackert, G. ; Sabel, M. ; Felsberg, J. ; Zacher, A. ; Kaulich, K. ; Hübschmann, D. ; Herold-Mende, C. ; von Deimling, A. ; Weller, M. ; Radlwimmer, B. ; Schlesner, M. ; Reifenberger, G. ; Höfer, T. ; Lichter, P.
Hellwig, M. ; Lauffer, M. C. ; Bockmayr, M. ; Spohn, M. ; Merk, D. J. ; Harrison, L. ; Ahlfeld, J. ; Kitowski, A. ; Neumann, J. E. ; Ohli, J. ; Holdhof, D. ; Niesen, J. ; Schoof, M. ; Kool, M. ; Kraus, C. ; Zweier, C. ; Holmberg, D. ; Schüller, U.
Buhl, J. L. ; Selt, F. ; Hielscher, T. ; Guiho, R. ; Ecker, J. ; Sahm, F. ; Ridinger, J. ; Riehl, D. ; Usta, D. ; Ismer, B. ; Sommerkamp, A. C. ; Martinez-Barbera, J. P. ; Wefers, A. K. ; Remke, M. ; Picard, D. ; Pusch, S. ; Gronych, J. ; Oehme, I. ; van Tilburg, C. M. ; Kool, M. ; Kuhn, D. ; Capper, D. ; von Deimling, A. ; Schuhmann, M. U. ; Herold-Mende, C. ; Korshunov, A. ; Brummer, T. ; Pfister, S. ; Jones, D. ; Witt, O. ; Milde, T.
Lee, J. C. ; Villanueva-Meyer, J. E. ; Ferris, S. P. ; Sloan, E. A. ; Hofmann, J. W. ; Hattab, E. M. ; Williams, B. J. ; Guo, H. ; Torkildson, J. ; Florez, A. ; Van Ziffle, J. ; Onodera, C. ; Grenert, J. P. ; Cho, S.-J. ; Horvai, A. E. ; Jones, D. ; Pfister, S. ; Koelsche, C. ; von Deimling, A. ; Korshunov, A. ; Perry, A. ; Solomon, D. A.
Korshunov, A. ; Casalini, B. ; Chavez, L. ; Hielscher, T. ; Sill, M. ; Ryzhova, M. ; Sharma, T. ; Schrimpf, D. ; Stichel, D. ; Capper, D. ; Reuss, D. E. ; Sturm, D. ; Absalyamova, O. ; Golanov, A. ; Lambo, S. ; Bewerunge-Hudler, M. ; Lichter, P. ; Herold-Mende, C. ; Wick, W. ; Pfister, S. ; Kool, M. ; Jones, D. ; von Deimling, A. ; Sahm, F.
Erkek, S. ; Johann, P. ; Finetti, M. A. ; Drosos, Y. ; Chou, H.-C. ; Zapatka, M. ; Sturm, D. ; Jones, D. ; Korshunov, A. ; Rhyzova, M. ; Wolf, S. ; Mallm, J.-P. ; Beck, K. ; Witt, O. ; Kulozik, A. E. ; Frühwald, M. C. ; Northcott, P. A. ; Korbel, J. O. ; Lichter, P. ; Eils, R. ; Gajjar, A. ; Roberts, C. W. M. ; Williamson, D. ; Hasselblatt, M. ; Chavez, L. ; Pfister, S. ; Kool, M.
Dietz, S. ; Lifshitz, A. ; Kazdal, D. ; Harms, A. ; Endris, V. ; Winter, H. ; Stenzinger, A. ; Warth, A. ; Sill, M. ; Tanay, A. ; Sültmann, H.
Uro-Coste, E. ; Masliah-Planchon, J. ; Siegfried, A. ; Blanluet, M. ; Lambo, S. ; Kool, M. ; Roujeau, T. ; Boetto, S. ; Palenzuela, G. ; Bertozzi, A.-I. ; Gambart, M. ; Coupier, I. ; Oliver-Petit, I. ; Golmard, L. ; Julia, S. ; Savagner, F. ; Mohand-Oumoussa, B. ; Tauziede-Espariat, A. ; Delisle, M.-B. ; Figarella-Branger, D. ; Bourdeaut, F. ; Rigau, V.
Korshunov, A. ; Sahm, F. ; Zheludkova, O. ; Golanov, A. ; Stichel, D. ; Schrimpf, D. ; Ryzhova, M. ; Potapov, A. ; Habel, A. ; Meyer, J. ; Lichter, P. ; Jones, D. ; von Deimling, A. ; Pfister, S. ; Kool, M.
Wirsching, H.-G. ; Tabatabai, G. ; Roelcke, U. ; Hottinger, A. F. ; Jörger, F. ; Schmid, A. ; Plasswilm, L. ; Schrimpf, D. ; Mancao, C. ; Capper, D. ; Conen, K. ; Hundsberger, T. ; Caparrotti, F. ; von Moos, R. ; Riklin, C. ; Felsberg, J. ; Roth, P. ; Jones, D. ; Pfister, S. ; Rushing, E. J. ; Abrey, L. ; Reifenberger, G. ; Held, L. ; von Deimling, A. ; Ochsenbein, A. ; Weller, M.
Rutkowski, S. ; Modena, P. ; Williamson, D. ; Kerl, K. ; Nysom, K. ; Pizer, B. ; Bartels, U. ; Puget, S. ; Doz, F. ; Michalski, A. ; von Hoff, K. ; Chevignard, M. ; Avula, S. ; Murray, M. J. ; Schönberger, S. ; Czech, T. ; Schouten-van Meeteren, A. Y. N. ; Kordes, U. ; Kramm, C. M. ; van Vuurden, D. G. ; Hulleman, E. ; Janssens, G. O. ; Solanki, G. A. ; van Veelen, M. C. ; Thomale, U. ; Schuhmann, M. U. ; Jones, C. ; Giangaspero, F. ; Figarella-Branger, D. ; Pietsch, T. ; Clifford, S. C. ; Pfister, S. ; Van Gool, S. W.
Ackermann, S. ; Cartolano, M. ; Hero, B. ; Welte, A. ; Kahlert, Y. ; Roderwieser, A. ; Bartenhagen, C. ; Walter, E. ; Gecht, J. ; Kerschke, L. ; Volland, R. ; Menon, R. ; Heuckmann, J. M. ; Gartlgruber, M. ; Hartlieb, S. ; Henrich, K.-O. ; Okonechnikov, K. ; Altmüller, J. ; Nürnberg, P. ; Lefever, S. ; de Wilde, B. ; Sand, F. ; Ikram, F. ; Rosswog, C. ; Fischer, J. ; Theissen, J. ; Hertwig, F. ; Singhi, A. D. ; Simon, T. ; Vogel, W. ; Perner, S. ; Krug, B. ; Schmidt, M. ; Rahmann, S. ; Achter, V. ; Lang, U. ; Vokuhl, C. ; Ortmann, M. ; Büttner, R. ; Eggert, A. ; Speleman, F. ; O'Sullivan, R. J. ; Thomas, R. K. ; Berthold, F. ; Vandesompele, J. ; Schramm, A. ; Westermann, F. ; Schulte, J. ; Peifer, M. ; Fischer, M.
Ratnaparkhe, M. ; Wong, J. K. L. ; Wei, P.-C. ; Hlevnjak, M. ; Kolb, T. ; Simovic, M. ; Haag, D. ; Paul, Y. ; Devens, F. ; Northcott, P. ; Jones, D. ; Kool, M. ; Jauch, A. ; Pastorczak, A. ; Mlynarski, W. ; Korshunov, A. ; Kumar, R. ; Downing, S. M. ; Pfister, S. ; Zapatka, M. ; McKinnon, P. J. ; Alt, F. W. ; Lichter, P. ; Ernst, A.
Tejada Neyra, M. A. ; Neuberger, U. ; Reinhardt, A. ; Brugnara, G. ; Bonekamp, D. ; Sill, M. ; Wick, A. ; Jones, D. ; Radbruch, A. ; Unterberg, A. ; Debus, J. ; Heiland, S. ; Schlemmer, H.-P. ; Herold-Mende, C. ; Pfister, S. ; von Deimling, A. ; Wick, W. ; Capper, D. ; Bendszus, M. ; Kickingereder, P.
Castel, D. ; Philippe, C. ; Kergrohen, T. ; Sill, M. ; Merlevede, J. ; Barret, E. ; Puget, S. ; Sainte-Rose, C. ; Kramm, C. M. ; Jones, C. ; Varlet, P. ; Pfister, S. ; Grill, J. ; Jones, D. ; Debily, M.-A.
Brabetz, S. ; Leary, S. E. S. ; Gröbner, S. ; Nakamoto, M. W. ; Şeker-Cin, H. ; Girard, E. J. ; Cole, B. ; Strand, A. D. ; Bloom, K. L. ; Hovestadt, V. ; Mack, N. ; Pakiam, F. ; Schwalm, B. ; Korshunov, A. ; Balasubramanian, G. P. ; Northcott, P. A. ; Pedro, K. D. ; Dey, J. ; Hansen, S. ; Ditzler, S. ; Lichter, P. ; Chavez, L. ; Jones, D. ; Koster, J. ; Pfister, S. ; Kool, M. ; Olson, J. M.
Sievers, P. ; Stichel, D. ; Hielscher, T. ; Schrimpf, D. ; Reinhardt, A. ; Wefers, A. K. ; Reuss, D. ; Jones, D. ; Bewerunge-Hudler, M. ; Hartmann, C. ; Baumgarten, P. ; Wirsching, H.-G. ; Winther-Kristensen, B. ; Brokinkel, B. ; Ketter, R. ; Idoate Gastearena, M. A. ; Lamszus, K. ; Seiz-Rosenhagen, M. ; Mawrin, C. ; Harter, P. ; Felsberg, J. ; Hänggi, D. ; Herold-Mende, C. ; Berghoff, A. S. ; Weller, M. ; Pfister, S. ; Wick, W. ; Reifenberger, G. ; Preusser, M. ; von Deimling, A. ; Sahm, F.
Hwang, E. I. ; Kool, M. ; Burger, P. C. ; Capper, D. ; Chavez, L. ; Brabetz, S. ; Williams-Hughes, C. ; Billups, C. ; Heier, L. ; Jaju, A. ; Michalski, J. ; Li, Y. ; Leary, S. ; Zhou, T. ; von Deimling, A. ; Jones, D. ; Fouladi, M. ; Pollack, I. F. ; Gajjar, A. ; Packer, R. J. ; Pfister, S. ; Olson, J. M.
Forget, A. ; Martignetti, L. ; Puget, S. ; Calzone, L. ; Brabetz, S. ; Picard, D. J. ; Montagud, A. ; Liva, S. ; Sta, A. ; Dingli, F. ; Arras, G. ; Rivera, J. ; Loew, D. ; Besnard, A. ; Lacombe, J. ; Pagès, M. ; Varlet, P. ; Dufour, C. ; Yu, H. ; Mercier, A. L. ; Indersie, E. ; Chivet, A. ; Leboucher, S. ; Sieber, L. ; Beccaria, K. ; Gombert, M. ; Meyer, F.-D. ; Qin, N. ; Bartl, J. ; Chavez, L. ; Okonechnikov, K. ; Sharma, T. ; Thatikonda, V. ; Bourdeaut, F. ; Pouponnot, C. ; Ramaswamy, V. ; Korshunov, A. ; Borkhardt, A. ; Reifenberger, G. ; Poullet, P. ; Taylor, M. D. ; Kool, M. ; Pfister, S. ; Kawauchi, D. ; Barillot, E. ; Remke, M. ; Ayrault, O.
Archer, T. C. ; Ehrenberger, T. ; Mundt, F. ; Gold, M. P. ; Krug, K. ; Mah, C. K. ; Mahoney, E. L. ; Daniel, C. J. ; LeNail, A. ; Ramamoorthy, D. ; Mertins, P. ; Mani, D. R. ; Zhang, H. ; Gillette, M. A. ; Clauser, K. ; Noble, M. ; Tang, L. C. ; Pierre-François, J. ; Silterra, J. ; Jensen, J. ; Tamayo, P. ; Korshunov, A. ; Pfister, S. ; Kool, M. ; Northcott, P. A. ; Sears, R. C. ; Lipton, J. O. ; Carr, S. A. ; Mesirov, J. P. ; Pomeroy, S. L. ; Fraenkel, E.
Koelsche, C. ; Hartmann, W. ; Schrimpf, D. ; Stichel, D. ; Jabar, S. ; Ranft, A. ; Reuss, D. E. ; Sahm, F. ; Jones, D. ; Bewerunge-Hudler, M. ; Trautmann, M. ; Klingebiel, T. ; Vokuhl, C. ; Gessler, M. ; Wardelmann, E. ; Petersen, I. ; Baumhoer, D. ; Flucke, U. ; Antonescu, C. ; Esteller, M. ; Fröhling, S. ; Kool, M. ; Pfister, S. ; Mechtersheimer, G. ; Dirksen, U. ; von Deimling, A.
Bächli, H. ; Ecker, J. ; van Tilburg, C. M. ; Sturm, D. ; Selt, F. ; Sahm, F. ; Koelsche, C. ; Grund, K. B. ; Sutter, C. ; Pietsch, T. ; Witt, H. ; Herold-Mende, C. ; von Deimling, A. ; Jones, D. ; Pfister, S. ; Witt, O. ; Milde, T.
Wegert, J. ; Vokuhl, C. ; Collord, G. ; Del Castillo Velasco-Herrera, M. ; Farndon, S. J. ; Guzzo, C. ; Jorgensen, M. ; Anderson, J. ; Slater, O. ; Duncan, C. ; Bausenwein, S. ; Streitenberger, H. ; Ziegler, B. ; Furtwängler, R. ; Graf, N. ; Stratton, M. R. ; Campbell, P. J. ; Jones, D. ; Koelsche, C. ; Pfister, S. ; Mifsud, W. ; Sebire, N. ; Sparber-Sauer, M. ; Koscielniak, E. ; Rosenwald, A. ; Gessler, M. ; Behjati, S.
Solomon, D. A. ; Korshunov, A. ; Sill, M. ; Jones, D. ; Kool, M. ; Pfister, S. ; Fan, X. ; Bannykh, S. ; Hu, J. ; Danielpour, M. ; Li, R. ; Johnston, J. ; Cham, E. ; Cooney, T. ; Sun, P. P. ; Oberheim Bush, N. A. ; McDermott, M. ; Van Ziffle, J. ; Onodera, C. ; Grenert, J. P. ; Bastian, B. C. ; Villanueva-Meyer, J. E. ; Pekmezci, M. ; Bollen, A. W. ; Perry, A.
Snuderl, M. ; Kannan, K. ; Pfaff, E. ; Wang, S. ; Stafford, J. M. ; Serrano, J. ; Heguy, A. ; Ray, K. ; Faustin, A. ; Aminova, O. ; Dolgalev, I. ; Stapleton, S. L. ; Zagzag, D. ; Chiriboga, L. ; Gardner, S. L. ; Wisoff, J. H. ; Golfinos, J. G. ; Capper, D. ; Hovestadt, V. ; Rosenblum, M. K. ; Placantonakis, D. G. ; LeBoeuf, S. E. ; Papagiannakopoulos, T. Y. ; Chavez, L. ; Ahsan, S. ; Eberhart, C. G. ; Pfister, S. ; Jones, D. ; Karajannis, M. A.
Sievers, P. ; Stichel, D. ; Schrimpf, D. ; Sahm, F. ; Koelsche, C. ; Reuss, D. E. ; Wefers, A. K. ; Reinhardt, A. ; Huang, K. ; Ebrahimi, A. ; Hou, Y. ; Pajtler, K. ; Pfister, S. ; Hasselblatt, M. ; Stummer, W. ; Schick, U. ; Hartmann, C. ; Hagel, C. ; Staszewski, O. ; Reifenberger, G. ; Beschorner, R. ; Coras, R. ; Keyvani, K. ; Kohlhof, P. ; Diomedi-Camassei, F. ; Herold-Mende, C. ; Giangaspero, F. ; Rushing, E. ; Giannini, C. ; Korshunov, A. ; Jones, D. ; von Deimling, A.
Shirahata, M. ; Ono, T. ; Stichel, D. ; Schrimpf, D. ; Reuss, D. E. ; Sahm, F. ; Koelsche, C. ; Wefers, A. ; Reinhardt, A. ; Huang, K. ; Sievers, P. ; Shimizu, H. ; Nanjo, H. ; Kobayashi, Y. ; Miyake, Y. ; Suzuki, T. ; Adachi, J.-I. ; Mishima, K. ; Sasaki, A. ; Nishikawa, R. ; Bewerunge-Hudler, M. ; Ryzhova, M. ; Absalyamova, O. ; Golanov, A. ; Sinn, P. ; Platten, M. ; Jungk, C. ; Winkler, F. ; Wick, A. ; Hänggi, D. ; Unterberg, A. ; Pfister, S. ; Jones, D. ; van den Bent, M. ; Hegi, M. ; French, P. ; Baumert, B. G. ; Stupp, R. ; Gorlia, T. ; Weller, M. ; Capper, D. ; Korshunov, A. ; Herold-Mende, C. ; Wick, W. ; Louis, D. N. ; von Deimling, A.
Pajtler, K. ; Wen, J. ; Sill, M. ; Lin, T. ; Orisme, W. ; Tang, B. ; Hübner, J.-M. ; Ramaswamy, V. ; Jia, S. ; Dalton, J. D. ; Haupfear, K. ; Rogers, H. A. ; Punchihewa, C. ; Lee, R. ; Easton, J. ; Wu, G. ; Ritzmann, T. A. ; Chapman, R. ; Chavez, L. ; Boop, F. A. ; Klimo, P. ; Sabin, N. D. ; Ogg, R. ; Mack, S. C. ; Freibaum, B. D. ; Kim, H. J. ; Witt, H. ; Jones, D. ; Vo, B. ; Gajjar, A. ; Pounds, S. ; Onar-Thomas, A. ; Roussel, M. F. ; Zhang, J. ; Taylor, J. P. ; Merchant, T. E. ; Grundy, R. ; Tatevossian, R. G. ; Taylor, M. D. ; Pfister, S. M. ; Korshunov, A. ; Kool, M. ; Ellison, D. W.
Koelsche, C. ; Mynarek, M. ; Schrimpf, D. ; Bertero, L. ; Serrano, J. ; Sahm, F. ; Reuss, D. E. ; Hou, Y. ; Baumhoer, D. ; Vokuhl, C. ; Flucke, U. ; Petersen, I. ; Brück, W. ; Rutkowski, S. ; Zambrano, S. C. ; Garcia Leon, J. L. ; Diaz Coronado, R. Y. ; Gessler, M. ; Tirado, O. M. ; Mora, J. ; Alonso, J. ; Garcia Del Muro, X. ; Esteller, M. ; Sturm, D. ; Ecker, J. ; Milde, T. ; Pfister, S. ; Korshunov, A. ; Snuderl, M. ; Mechtersheimer, G. ; Schüller, U. ; Jones, D. ; von Deimling, A.
Deng, M. Y. ; Sill, M. ; Chiang, J. ; Schittenhelm, J. ; Ebinger, M. ; Schuhmann, M. U. ; Monoranu, C.-M. ; Milde, T. ; Wittmann, A. ; Hartmann, C. ; Sommer, C. ; Paulus, W. ; Gärtner, J. ; Brück, W. ; Rüdiger, T. ; Leipold, A. ; Jaunmuktane, Z. ; Brandner, S. ; Giangaspero, F. ; Nozza, P. ; Mora, J. ; Morales la Madrid, A. ; Cruz Martinez, O. ; Hansford, J. R. ; Pietsch, T. ; Tietze, A. ; Hernáiz-Driever, P. ; Stoler, I. ; Capper, D. ; Korshunov, A. ; Ellison, D. W. ; von Deimling, A. ; Pfister, S. ; Sahm, F. ; Jones, D.
Capper, D. ; Stichel, D. ; Sahm, F. ; Jones, D. ; Schrimpf, D. ; Sill, M. ; Schmid, S. ; Hovestadt, V. ; Reuss, D. E. ; Koelsche, C. ; Reinhardt, A. ; Wefers, A. K. ; Huang, K. ; Sievers, P. ; Ebrahimi, A. ; Schöler, A. ; Teichmann, D. ; Koch, A. ; Hänggi, D. ; Unterberg, A. ; Platten, M. ; Wick, W. ; Witt, O. ; Milde, T. ; Korshunov, A. ; Pfister, S. ; von Deimling, A.
Capper, D. ; Engel, N. W. ; Stichel, D. ; Lechner, M. ; Glöss, S. ; Schmid, S. ; Koelsche, C. ; Schrimpf, D. ; Niesen, J. ; Wefers, A. K. ; Jones, D. ; Sill, M. ; Weigert, O. ; Ligon, K. L. ; Olar, A. ; Koch, A. ; Forster, M. ; Moran, S. ; Tirado, O. M. ; Sáinz-Jaspeado, M. ; Mora, J. ; Esteller, M. ; Alonso, J. ; Del Muro, X. G. ; Paulus, W. ; Felsberg, J. ; Reifenberger, G. ; Glatzel, M. ; Frank, S. ; Monoranu, C. M. ; Lund, V. J. ; von Deimling, A. ; Pfister, S. ; Buslei, R. ; Ribbat-Idel, J. ; Perner, S. ; Gudziol, V. ; Meinhardt, M. ; Schüller, U.
Akgül, S. ; Li, Y. ; Zheng, S. ; Kool, M. ; Treisman, D. M. ; Li, C. ; Wang, Y. ; Gröbner, S. ; Ikenoue, T. ; Shen, Y. ; Camelo-Piragua, S. ; Tomasek, G. ; Stark, S. ; Guduguntla, V. ; Gusella, J. F. ; Guan, K.-L. ; Pfister, S. ; Verhaak, R. G. W. ; Zhu, Y.
Robinson, G. W. ; Rudneva, V. A. ; Buchhalter, I. ; Billups, C. A. ; Waszak, S. M. ; Smith, K. S. ; Bowers, D. C. ; Bendel, A. ; Fisher, P. G. ; Partap, S. ; Crawford, J. R. ; Hassall, T. ; Indelicato, D. J. ; Boop, F. ; Klimo, P. ; Sabin, N. D. ; Patay, Z. ; Merchant, T. E. ; Stewart, C. F. ; Orr, B. A. ; Korbel, J. O. ; Jones, D. ; Sharma, T. ; Lichter, P. ; Kool, M. ; Korshunov, A. ; Pfister, S. ; Gilbertson, R. J. ; Sanders, R. P. ; Onar-Thomas, A. ; Ellison, D. W. ; Gajjar, A. ; Northcott, P. A.
Gozdecka, M. ; Meduri, E. ; Mazan, M. ; Tzelepis, K. ; Dudek, M. ; Knights, A. J. ; Pardo, M. ; Yu, L. ; Choudhary, J. S. ; Metzakopian, E. ; Iyer, V. ; Yun, H. ; Park, N. ; Varela, I. ; Bautista, R. ; Collord, G. ; Dovey, O. ; Garyfallos, D. A. ; De Braekeleer, E. ; Kondo, S. ; Cooper, J. ; Göttgens, B. ; Bullinger, L. ; Northcott, P. A. ; Adams, D. ; Vassiliou, G. S. ; Huntly, B. J. P.
Pfaff, E. ; Kessler, T. ; Balasubramanian, G. P. ; Berberich, A. ; Schrimpf, D. ; Wick, A. ; Debus, J. ; Unterberg, A. ; Bendszus, M. ; Herold-Mende, C. ; Capper, D. ; Schenkel, I. ; Eisenmenger, A. ; Dettmer, S. ; Brors, B. ; Platten, M. ; Pfister, S. ; von Deimling, A. ; Jones, D. ; Wick, W. ; Sahm, F.
Kickingereder, P. ; Neuberger, U. ; Bonekamp, D. ; Piechotta, P. L. ; Götz, M. ; Wick, A. ; Sill, M. ; Kratz, A. ; Shinohara, R. T. ; Jones, D. ; Radbruch, A. ; Muschelli, J. ; Unterberg, A. ; Debus, J. ; Schlemmer, H.-P. ; Herold-Mende, C. ; Pfister, S. ; von Deimling, A. ; Wick, W. ; Capper, D. ; Maier-Hein, K. ; Bendszus, M.
Mackay, A. ; Burford, A. ; Molinari, V. ; Jones, D. ; Izquierdo, E. ; Brouwer-Visser, J. ; Giangaspero, F. ; Haberler, C. ; Pietsch, T. ; Jacques, T. S. ; Figarella-Branger, D. ; Rodriguez, D. ; Morgan, P. S. ; Raman, P. ; Waanders, A. J. ; Resnick, A. C. ; Massimino, M. ; Garrè, M. L. ; Smith, H. ; Capper, D. ; Pfister, S. ; Würdinger, T. ; Tam, R. ; Garcia, J. ; Thakur, M. D. ; Vassal, G. ; Grill, J. ; Jaspan, T. ; Varlet, P. ; Jones, C.
Apps, J. R. ; Carreno, G. ; Gonzalez-Meljem, J. M. ; Haston, S. ; Guiho, R. ; Cooper, J. E. ; Manshaei, S. ; Jani, N. ; Hölsken, A. ; Pettorini, B. ; Beynon, R. J. ; Simpson, D. M. ; Fraser, H. C. ; Hong, Y. ; Hallang, S. ; Stone, T. J. ; Virasami, A. ; Donson, A. M. ; Jones, D. ; Aquilina, K. ; Spoudeas, H. ; Joshi, A. R. ; Grundy, R. ; Storer, L. C. D. ; Korbonits, M. ; Hilton, D. A. ; Tossell, K. ; Thavaraj, S. ; Ungless, M. A. ; Gil, J. ; Buslei, R. ; Hankinson, T. ; Hargrave, D. ; Goding, C. ; Andoniadou, C. L. ; Brogan, P. ; Jacques, T. S. ; Williams, H. J. ; Martinez-Barbera, J. P.
Johann, P. ; Bens, S. ; Oyen, F. ; Wagener, R. ; Giannini, C. ; Perry, A. ; Raisanen, J. M. ; Reis, G. F. ; Nobusawa, S. ; Arita, K. ; Felsberg, J. ; Reifenberger, G. ; Agaimy, A. ; Buslei, R. ; Capper, D. ; Pfister, S. ; Schneppenheim, R. ; Siebert, R. ; Frühwald, M. C. ; Paulus, W. ; Kool, M. ; Hasselblatt, M.
Merk, D. J. ; Ohli, J. ; Merk, N. D. ; Thatikonda, V. ; Morrissy, S. ; Schoof, M. ; Schmid, S. N. ; Harrison, L. ; Filser, S. ; Ahlfeld, J. ; Erkek, S. ; Raithatha, K. ; Andreska, T. ; Weißhaar, M. ; Launspach, M. ; Neumann, J. E. ; Shakarami, M. ; Plenker, D. ; Marra, M. A. ; Li, Y. ; Mungall, A. J. ; Moore, R. A. ; Ma, Y. ; Jones, S. J. M. ; Lutz, B. ; Ertl-Wagner, B. ; Rossi, A. ; Wagener, R. ; Siebert, R. ; Jung, A. ; Eberhart, C. G. ; Lach, B. ; Sendtner, M. ; Pfister, S. ; Taylor, M. D. ; Chavez, L. ; Kool, M. ; Schüller, U.
Iorgulescu, J. B. ; Van Ziffle, J. ; Stevers, M. ; Grenert, J. P. ; Bastian, B. C. ; Chavez, L. ; Stichel, D. ; Buchhalter, I. ; Samuel, D. ; Nicolaides, T. ; Banerjee, A. ; Mueller, S. ; Gupta, N. ; Tihan, T. ; Bollen, A. W. ; Northcott, P. A. ; Kool, M. ; Pfister, S. ; Korshunov, A. ; Perry, A. ; Solomon, D. A.
Gómez, S. ; Garrido-Garcia, A. ; Garcia-Gerique, L. ; Lemos, I. ; Suñol, M. ; de Torres, C. ; Kulis, M. ; Pérez-Jaume, S. ; Carcaboso, Á. M. ; Luu, B. ; Kieran, M. W. ; Jabado, N. ; Kozlenkov, A. ; Dracheva, S. ; Ramaswamy, V. ; Hovestadt, V. ; Johann, P. ; Jones, D. ; Pfister, S. ; Morales La Madrid, A. ; Cruz, O. ; Taylor, M. D. ; Martin-Subero, J.-I. ; Mora, J. ; Lavarino, C.
Infante, P. ; Faedda, R. ; Bernardi, F. ; Bufalieri, F. ; Lospinoso Severini, L. ; Alfonsi, R. ; Mazzà, D. ; Siler, M. ; Coni, S. ; Po, A. ; Petroni, M. ; Ferretti, E. ; Mori, M. ; De Smaele, E. ; Canettieri, G. ; Capalbo, C. ; Maroder, M. ; Screpanti, I. ; Kool, M. ; Pfister, S. ; Guardavaccaro, D. ; Gulino, A. ; Di Marcotullio, L.
Kessler, T. ; Sahm, F. ; Sadik, A. ; Stichel, D. ; Hertenstein, A. ; Reifenberger, G. ; Zacher, A. ; Sabel, M. ; Tabatabai, G. ; Steinbach, J. ; Sure, U. ; Krex, D. ; Grosu, A.-L. ; Bewerunge-Hudler, M. ; Jones, D. ; Pfister, S. ; Weller, M. ; Opitz, C. ; Bendszus, M. ; von Deimling, A. ; Platten, M. ; Wick, W.
Jones, D. ; Kieran, M. W. ; Bouffet, E. ; Alexandrescu, S. ; Bandopadhayay, P. ; Bornhorst, M. ; Ellison, D. ; Fangusaro, J. ; Fisher, M. J. ; Foreman, N. ; Fouladi, M. ; Hargrave, D. ; Hawkins, C. ; Jabado, N. ; Massimino, M. ; Mueller, S. ; Perilongo, G. ; Schouten van Meeteren, A. Y. N. ; Tabori, U. ; Warren, K. ; Waanders, A. J. ; Walker, D. ; Weiss, W. ; Witt, O. ; Wright, K. ; Zhu, Y. ; Bowers, D. C. ; Pfister, S. ; Packer, R. J.
Beier, D. ; Proescholdt, M. ; Reinert, C. ; Pietsch, T. ; Jones, D. ; Pfister, S. ; Hattingen, E. ; Seidel, C. ; Dirven, L. ; Luerding, R. ; Reijneveld, J. ; Warmuth-Metz, M. ; Bonsanto, M. ; Bremer, M. ; Combs, S. E. ; Rieken, S. ; Herrlinger, U. ; Kuntze, H. ; Mayer-Steinacker, R. ; Moskopp, D. ; Schneider, T. ; Beringer, A. ; Schlegel, U. ; Stummer, W. ; Welker, H. ; Weyerbrock, A. ; Paulsen, F. ; Rutkowski, S. ; Weller, M. ; Wick, W. ; Kortmann, R.-D. ; Bogdahn, U. ; Hau, P.
van Tilburg, C. M. ; Selt, F. ; Sahm, F. ; Bächli, H. ; Pfister, S. ; Witt, O. ; Milde, T.
Nemes, K. ; Bens, S. ; Bourdeaut, F. ; Johann, P. ; Kordes, U. ; Siebert, S. ; Frühwald
Deeg, K. ; Chung, I. ; Poos, A. M. ; Braun, D. M. ; Korshunov, A. ; Oswald, M. ; Kepper, N. ; Bender, S. ; Castel, D. ; Lichter, P. ; Grill, J. ; Pfister, S. M. ; König, R. ; Jones, D. T. W. ; Rippe, K.
Bartenhagen, C. ; Fischer, U. ; Korn, K. ; Pfister, S. ; Gombert, M. ; Chen, C. ; Okpanyi, V. ; Hauer, J. ; Rinaldi, A. ; Bourquin, J.-P. ; Eckert, C. ; Hu, J. ; Ensser, A. ; Dugas, M. ; Borkhardt, A.
Wu, X. ; Zhang, L.-S. ; Toombs, J. ; Kuo, Y.-C. ; Piazza, J. T. ; Tuladhar, R. ; Barrett, Q. ; Fan, C.-W. ; Zhang, X. ; Walensky, L. D. ; Kool, M. ; Cheng, S. Y. ; Brekken, R. ; Opferman, J. T. ; Green, D. R. ; Moldoveanu, T. ; Lum, L.
Schmidt, C. ; Schubert, N. A. ; Brabetz, S. ; Mack, N. ; Schwalm, B. ; Chan, J. A. ; Selt, F. ; Herold-Mende, C. ; Witt, O. ; Milde, T. ; Pfister, S. ; Korshunov, A. ; Kool, M.
Andreiuolo, F. ; Le Teuff, G. ; Bayar, M. A. ; Kilday, J.-P. ; Pietsch, T. ; von Bueren, A. O. ; Witt, H. ; Korshunov, A. ; Modena, P. ; Pfister, S. ; Pagès, M. ; Castel, D. ; Giangaspero, F. ; Chimelli, L. ; Varlet, P. ; Rutkowski, S. ; Frappaz, D. ; Massimino, M. ; Grundy, R. ; Grill, J. ; BIOMECA, S. E. B. W. G.
Koelsche, C. ; Schrimpf, D. ; Tharun, L. ; Roth, E. ; Sturm, D. ; Jones, D. ; Renker, E.-K. ; Sill, M. ; Baude, A. ; Sahm, F. ; Capper, D. ; Bewerunge-Hudler, M. ; Hartmann, W. ; Kulozik, A. E. ; Petersen, I. ; Flucke, U. ; Schreuder, H. W. B. ; Büttner, R. ; Weber, M.-A. ; Schirmacher, P. ; Plass, C. ; Pfister, S. ; von Deimling, A. ; Mechtersheimer, G.
Senfter, D. ; Erkan, E. P. ; Özer, E. ; Jungwirth, G. ; Madlener, S. ; Kool, M. ; Ströbel, T. ; Saydam, N. ; Saydam, O.
Panwalkar, P. ; Clark, J. ; Ramaswamy, V. ; Hawes, D. ; Yang, F. ; Dunham, C. ; Yip, S. ; Hukin, J. ; Sun, Y. ; Schipper, M. J. ; Chavez, L. ; Margol, A. ; Pekmezci, M. ; Chung, C. ; Banda, A. ; Bayliss, J. M. ; Curry, S. J. ; Santi, M. ; Rodriguez, F. J. ; Snuderl, M. ; Karajannis, M. A. ; Saratsis, A. M. ; Horbinski, C. M. ; Carret, A.-S. ; Wilson, B. ; Johnston, D. ; Lafay-Cousin, L. ; Zelcer, S. ; Eisenstat, D. ; Silva, M. ; Scheinemann, K. ; Jabado, N. ; McNeely, P. D. ; Kool, M. ; Pfister, S. ; Taylor, M. D. ; Hawkins, C. ; Korshunov, A. ; Judkins, A. R. ; Venneti, S.
Moreno, N. ; Holsten, T. ; Mertins, J. ; Zhogbi, A. ; Johann, P. ; Kool, M. ; Meisterernst, M. ; Kerl, K.
Neumann, J. E. ; Wefers, A. K. ; Lambo, S. ; Bianchi, E. ; Bockstaller, M. ; Dorostkar, M. M. ; Meister, V. ; Schindler, P. ; Korshunov, A. ; von Hoff, K. ; Nowak, J. ; Warmuth-Metz, M. ; Schneider, M. R. ; Renner-Müller, I. ; Merk, D. J. ; Shakarami, M. ; Sharma, T. ; Chavez, L. ; Glass, R. ; Chan, J. A. ; Taketo, M. M. ; Neumann, P. ; Kool, M. ; Schüller, U.
Ratnaparkhe, M. ; Hlevnjak, M. ; Kolb, T. ; Jauch, A. ; Maass, K. ; Devens, F. ; Rode, A. ; Hovestadt, V. ; Korshunov, A. ; Pastorczak, A. ; Mlynarski, W. ; Sungalee, S. ; Korbel, J. ; Hoell, J. ; Fischer, U. ; Milde, T. ; Kramm, C. ; Nathrath, M. ; Chrzanowska, K. ; Tausch, E. ; Takagi, M. ; Taga, T. ; Constantini, S. ; Loeffen, J. ; Meijerink, J. ; Zielen, S. ; Gohring, G. ; Schlegelberger, B. ; Maass, E. ; Siebert, R. ; Kunz, J. ; Kulozik, A. E. ; Worst, B. ; Jones, D. ; Pfister, S. ; Zapatka, M. ; Lichter, P. ; Ernst, A.
Huang, Z. ; Jones, D. ; Wu, Y. ; Lichter, P. ; Zapatka, M.
Korshunov, A. ; Schrimpf, D. ; Ryzhova, M. ; Sturm, D. ; Chavez, L. ; Hovestadt, V. ; Sharma, T. ; Habel, A. ; Burford, A. ; Jones, C. ; Zheludkova, O. ; Kumirova, E. ; Kramm, C. M. ; Golanov, A. ; Capper, D. ; von Deimling, A. ; Pfister, S. ; Jones, D.
Gojo, J. ; Lötsch, D. ; Spiegl-Kreinecker, S. ; Pajtler, K. ; Neumayer, K. ; Korbel, P. ; Araki, A. ; Brandstetter, A. ; Mohr, T. ; Hovestadt, V. ; Chavez, L. ; Kirchhofer, D. ; Ricken, G. ; Stefanits, H. ; Korshunov, A. ; Pfister, S. ; Dieckmann, K. ; Azizi, A. A. ; Czech, T. ; Filipits, M. ; Kool, M. ; Peyrl, A. ; Slavc, I. ; Berger, W. ; Haberler, C.
Bingel, C. ; Koeneke, E. ; Ridinger, J. ; Bittmann, A. ; Sill, M. ; Peterziel, H. ; Wrobel, J. ; Rettig, I. ; Milde, T. ; Fernekorn, U. ; Weise, F. ; Schober, A. ; Witt, O. ; Oehme, I.
Sahm, F. ; Toprak, U. ; Hübschmann, D. ; Kleinheinz, K. ; Buchhalter, I. ; Sill, M. ; Stichel, D. ; Schick, M. ; Bewerunge-Hudler, M. ; Schrimpf, D. ; Zadeh, G. ; Aldape, K. ; Herold-Mende, C. ; Beck, K. ; Staszewski, O. ; Prinz, M. ; Harosh, C. B. ; Eils, R. ; Sturm, D. ; Jones, D. ; Pfister, S. ; Paulus, W. ; Ram, Z. ; Schlesner, M. ; Grossman, R. ; von Deimling, A.
Pearson, A. D. J. ; Pfister, S. ; Baruchel, A. ; Bourquin, J.-P. ; Casanova, M. ; Chesler, L. ; Doz, F. ; Eggert, A. ; Geoerger, B. ; Jones, D. ; Kearns, P. R. ; Molenaar, J. J. ; Morland, B. ; Schleiermacher, G. ; Schulte, J. ; Vormoor, J. ; Marshall, L. V. ; Zwaan, C. M. ; Vassal, G. ; Consortium, E. a. B. C. o. t. I. T. f. C. w. C. E.
Johann, P. ; Hovestadt, V. ; Thomas, C. ; Jeibmann, A. ; Heß, K. ; Bens, S. ; Oyen, F. ; Hawkins, C. ; Pierson, C. R. ; Aldape, K. ; Kim, S.-P. ; Widing, E. ; Sumerauer, D. ; Hauser, P. ; van Landeghem, F. ; Ryzhova, M. ; Korshunov, A. ; Capper, D. ; Jones, D. ; Pfister, S. ; Schneppenheim, R. ; Siebert, R. ; Paulus, W. ; Frühwald, M. C. ; Kool, M. ; Hasselblatt, M.
Selt, F. ; Hohloch, J. ; Hielscher, T. ; Sahm, F. ; Capper, D. ; Korshunov, A. ; Usta, D. ; Brabetz, S. ; Ridinger, J. ; Ecker, J. ; Oehme, I. ; Gronych, J. ; Marquardt, V. ; Pauck, D. ; Bächli, H. ; Stiles, C. D. ; von Deimling, A. ; Remke, M. ; Schuhmann, M. U. ; Pfister, S. ; Brummer, T. ; Jones, D. ; Witt, O. ; Milde, T.
Sahm, F. ; Schrimpf, D. ; Stichel, D. ; Jones, D. ; Hielscher, T. ; Schefzyk, S. ; Okonechnikov, K. ; Koelsche, C. ; Reuss, D. E. ; Capper, D. ; Sturm, D. ; Wirsching, H.-G. ; Berghoff, A. S. ; Baumgarten, P. ; Kratz, A. ; Huang, K. ; Wefers, A. K. ; Hovestadt, V. ; Sill, M. ; Ellis, H. P. ; Kurian, K. M. ; Okuducu, A. F. ; Jungk, C. ; Drueschler, K. ; Schick, M. ; Bewerunge-Hudler, M. ; Mawrin, C. ; Seiz-Rosenhagen, M. ; Ketter, R. ; Simon, M. ; Westphal, M. ; Lamszus, K. ; Becker, A. ; Koch, A. ; Schittenhelm, J. ; Rushing, E. J. ; Collins, V. P. ; Brehmer, S. ; Chavez, L. ; Platten, M. ; Hänggi, D. ; Unterberg, A. ; Paulus, W. ; Wick, W. ; Pfister, S. ; Mittelbronn, M. ; Preusser, M. ; Herold-Mende, C. ; Weller, M. ; von Deimling, A.
Packer, R. J. ; Pfister, S. ; Bouffet, E. ; Avery, R. ; Bandopadhayay, P. ; Bornhorst, M. ; Bowers, D. C. ; Ellison, D. ; Fangusaro, J. ; Foreman, N. ; Fouladi, M. ; Gajjar, A. ; Haas-Kogan, D. ; Hawkins, C. ; Ho, C.-Y. ; Hwang, E. ; Jabado, N. ; Kilburn, L. B. ; Lassaletta, A. ; Ligon, K. L. ; Massimino, M. ; Meeteren, S.-v. ; Mueller, S. ; Nicolaides, T. ; Perilongo, G. ; Tabori, U. ; Vezina, G. ; Warren, K. ; Witt, O. ; Zhu, Y. ; Jones, D. ; Kieran, M.
Jones, C. ; Karajannis, M. A. ; Jones, D. ; Kieran, M. W. ; Monje, M. ; Baker, S. J. ; Becher, O. J. ; Cho, Y.-J. ; Gupta, N. ; Hawkins, C. ; Hargrave, D. ; Haas-Kogan, D. A. ; Jabado, N. ; Li, X.-N. ; Mueller, S. ; Nicolaides, T. ; Packer, R. J. ; Persson, A. I. ; Phillips, J. J. ; Simonds, E. F. ; Stafford, J. M. ; Tang, Y. ; Pfister, S. ; Weiss, W. A.
Kratz, C. P. ; Achatz, M. I. ; Brugières, L. ; Frebourg, T. ; Garber, J. E. ; Greer, M. C. ; Hansford, J. R. ; Janeway, K. A. ; Kohlmann, W. K. ; McGee, R. ; Mullighan, C. G. ; Onel, K. ; Pajtler, K. ; Pfister, S. ; Savage, S. A. ; Schiffman, J. D. ; Schneider, K. A. ; Strong, L. C. ; Evans, D. G. R. ; Wasserman, J. D. ; Villani, A. ; Malkin, D.
Mynarek, M. ; Pizer, B. ; Dufour, C. ; van Vuurden, D. ; Garami, M. ; Massimino, M. ; Fangusaro, J. ; Davidson, T. ; Gil-da-Costa, M. J. ; Sterba, J. ; Benesch, M. ; Gerber, N. ; Juhnke, B. O. ; Kwiecien, R. ; Pietsch, T. ; Kool, M. ; Clifford, S. ; Ellison, D. W. ; Giangaspero, F. ; Wesseling, P. ; Gilles, F. ; Gottardo, N. ; Finlay, J. L. ; Rutkowski, S. ; von Hoff, K.
Vragniau, C. ; Hübner, J.-M. ; Beidler, P. ; Gil, S. ; Saydaminova, K. ; Lu, Z.-Z. ; Yumul, R. ; Wang, H. ; Richter, M. ; Sova, P. ; Drescher, C. ; Fender, P. ; Lieber, A.
Janssens, G. O. ; Gandola, L. ; Bolle, S. ; Mandeville, H. ; Ramos-Albiac, M. ; van Beek, K. ; Benghiat, H. ; Hoeben, B. ; Morales La Madrid, A. ; Kortmann, R.-D. ; Hargrave, D. ; Menten, J. ; Pecori, E. ; Biassoni, V. ; von Bueren, A. O. ; van Vuurden, D. G. ; Massimino, M. ; Sturm, D. ; Peters, M. ; Kramm, C. M.
Feng, W. ; Kawauchi, D. ; Körkel-Qu, H. ; Deng, H. ; Serger, E. ; Sieber, L. ; Lieberman, J. A. ; Jimeno-González, S. ; Lambo, S. ; Hanna, B. ; Harim, Y. ; Jansen, M. ; Neuerburg, A. ; Friesen, O. ; Zuckermann, M. ; Rajendran, V. ; Gronych, J. ; Ayrault, O. ; Korshunov, A. ; Jones, D. ; Kool, M. ; Northcott, P. A. ; Lichter, P. ; Cortés-Ledesma, F. ; Pfister, S. ; Liu, H.
Sahm, F. ; Korshunov, A. ; Schrimpf, D. ; Stichel, D. ; Jones, D. ; Capper, D. ; Kölsche, C. ; Reuss, D. ; Kratz, A. ; Huang, K. ; Wefers, A. K. ; Schick, M. ; Bewerunge-Hudler, M. ; Mittelbronn, M. ; Platten, M. ; Hänggi, D. ; Jeibmann, A. ; Unterberg, A. ; Herold-Mende, C. ; Pfister, S. ; Brandner, S. ; Wick, W. ; von Deimling, A.
Pajtler, K. ; Mack, S. C. ; Ramaswamy, V. ; Smith, C. A. ; Witt, H. ; Smith, A. ; Hansford, J. R. ; von Hoff, K. ; Wright, K. D. ; Hwang, E. ; Frappaz, D. ; Kanemura, Y. ; Massimino, M. ; Faure-Conter, C. ; Modena, P. ; Tabori, U. ; Warren, K. E. ; Holland, E. C. ; Ichimura, K. ; Giangaspero, F. ; Castel, D. ; von Deimling, A. ; Kool, M. ; Dirks, P. B. ; Grundy, R. G. ; Foreman, N. K. ; Gajjar, A. ; Korshunov, A. ; Finlay, J. ; Gilbertson, R. J. ; Ellison, D. W. ; Aldape, K. D. ; Merchant, T. E. ; Bouffet, E. ; Pfister, S. ; Taylor, M. D.
Gunkel, M. ; Chung, I. ; Wörz, S. ; Deeg, K. ; Simon, R. ; Sauter, G. ; Jones, D. ; Korshunov, A. ; Rohr, K. ; Erfle, H. ; Rippe, K.
Weischenfeldt, J. ; Dubash, T. D. ; Drainas, A. P. ; Mardin, B. R. ; Chen, Y. ; Stütz, A. M. ; Waszak, S. M. ; Bosco, G. ; Halvorsen, A. R. ; Raeder, B. ; Efthymiopoulos, T. ; Erkek, S. ; Siegl, C. ; Brenner, H. ; Brustugun, O. T. ; Dieter, S. ; Northcott, P. A. ; Petersen, I. ; Pfister, S. ; Schneider, M. ; Solberg, S. K. ; Thunissen, E. ; Weichert, W. ; Zichner, T. ; Thomas, R. ; Peifer, M. ; Helland, A. ; Ball, C. ; Jechlinger, M. ; Sotillo, R. ; Glimm, H. ; Korbel, J. O.
TP53 codon 72 polymorphism may predict early tumour progression in paediatric pilocytic astrocytoma.
Mascelli, S. ; Nozza, P. ; Jones, D. ; Colin, C. ; Pistorio, A. ; Milanaccio, C. ; Ravegnani, M. ; Consales, A. ; Witt, O. ; Morana, G. ; Cama, A. ; Capra, V. ; Biassoni, R. ; Pfister, S. ; Figarella-Branger, D. ; Garrè, M. L. ; Raso, A.
von Bueren, A. O. ; Kortmann, R.-D. ; von Hoff, K. ; Friedrich, C. ; Mynarek, M. ; Müller, K. ; Goschzik, T. ; Zur Mühlen, A. ; Gerber, N. ; Warmuth-Metz, M. ; Soerensen, N. ; Deinlein, F. ; Benesch, M. ; Zwiener, I. ; Kwiecien, R. ; Faldum, A. ; Bode, U. ; Fleischhack, G. ; Hovestadt, V. ; Kool, M. ; Jones, D. ; Northcott, P. ; Kuehl, J. ; Pfister, S. ; Pietsch, T. ; Rutkowski, S.
Tzaridis, T. ; Milde, T. ; Pajtler, K. ; Bender, S. ; Jones, D. ; Müller, S. ; Wittmann, A. ; Schlotter, M. ; Kulozik, A. E. ; Lichter, P. ; Peter Collins, V. ; Witt, O. ; Kool, M. ; Korshunov, A. ; Pfister, S. ; Witt, H.
Thomas, C. ; Sill, M. ; Ruland, V. ; Witten, A. ; Hartung, S. ; Kordes, U. ; Jeibmann, A. ; Beschorner, R. ; Keyvani, K. ; Bergmann, M. ; Mittelbronn, M. ; Pietsch, T. ; Felsberg, J. ; Monoranu, C. M. ; Varlet, P. ; Hauser, P. ; Olar, A. ; Grundy, R. G. ; Wolff, J. E. ; Korshunov, A. ; Jones, D. ; Bewerunge-Hudler, M. ; Hovestadt, V. ; von Deimling, A. ; Pfister, S. ; Paulus, W. ; Capper, D. ; Hasselblatt, M.
Selt, F. ; Deiß, A. ; Korshunov, A. ; Capper, D. ; Witt, H. ; van Tilburg, C. M. ; Jones, D. ; Witt, R. ; Sahm, F. ; Reuss, D. ; Kölsche, C. ; Ecker, J. ; Oehme, I. ; Hielscher, T. ; von Deimling, A. ; Kulozik, A. E. ; Pfister, S. ; Witt, O. ; Milde, T.
Sahm, F. ; Schrimpf, D. ; Jones, D. ; Meyer, J. ; Kratz, A. ; Reuss, D. ; Capper, D. ; Koelsche, C. ; Korshunov, A. ; Wiestler, B. P. O. ; Buchhalter, I. ; Milde, T. ; Selt, F. ; Sturm, D. ; Kool, M. ; Hummel, M. ; Bewerunge-Hudler, M. ; Mawrin, C. ; Schüller, U. ; Jungk, C. ; Wick, A. ; Witt, O. ; Platten, M. ; Herold-Mende, C. ; Unterberg, A. ; Pfister, S. ; Wick, W. ; von Deimling, A.
Röhrich, M. ; Koelsche, C. ; Schrimpf, D. ; Capper, D. ; Sahm, F. ; Kratz, A. ; Reuss, J. ; Hovestadt, V. ; Jones, D. ; Bewerunge-Hudler, M. ; Becker, A. ; Weis, J. ; Mawrin, C. ; Mittelbronn, M. ; Perry, A. ; Mautner, V.-F. ; Mechtersheimer, G. ; Hartmann, C. ; Okuducu, A. F. ; Arp, M. ; Seiz-Rosenhagen, M. ; Hänggi, D. ; Heim, S. ; Paulus, W. ; Schittenhelm, J. ; Ahmadi, R. ; Herold-Mende, C. ; Unterberg, A. ; Pfister, S. ; von Deimling, A. ; Reuss, D. E.
Rivera, B. ; Gayden, T. ; Carrot-Zhang, J. ; Nadaf, J. ; Boshari, T. ; Faury, D. ; Zeinieh, M. ; Blanc, R. ; Burk, D. L. ; Fahiminiya, S. ; Bareke, E. ; Schüller, U. ; Monoranu, C. M. ; Sträter, R. ; Kerl, K. ; Niederstadt, T. ; Kurlemann, G. ; Ellezam, B. ; Michalak, Z. ; Thom, M. ; Lockhart, P. J. ; Leventer, R. J. ; Ohm, M. ; MacGregor, D. ; Jones, D. ; Karamchandani, J. ; Greenwood, C. M. T. ; Berghuis, A. M. ; Bens, S. ; Siebert, R. ; Zakrzewska, M. ; Liberski, P. P. ; Zakrzewski, K. ; Sisodiya, S. M. ; Paulus, W. ; Albrecht, S. ; Hasselblatt, M. ; Jabado, N. ; Foulkes, W. D. ; Majewski, J.
Ramaswamy, V. ; Remke, M. ; Bouffet, E. ; Bailey, S. ; Clifford, S. C. ; Doz, F. ; Kool, M. ; Dufour, C. ; Vassal, G. ; Milde, T. ; Witt, O. ; von Hoff, K. ; Pietsch, T. ; Northcott, P. A. ; Gajjar, A. ; Robinson, G. W. ; Padovani, L. ; André, N. ; Massimino, M. ; Pizer, B. ; Packer, R. ; Rutkowski, S. ; Pfister, S. ; Taylor, M. D. ; Pomeroy, S. L.
Pei, Y. ; Liu, K.-W. ; Wang, J. ; Garancher, A. ; Tao, R. ; Esparza, L. A. ; Maier, D. L. ; Udaka, Y. T. ; Murad, N. ; Morrissy, S. ; Seker-Cin, H. ; Brabetz, S. ; Qi, L. ; Kogiso, M. ; Schubert, S. ; Olson, J. M. ; Cho, Y.-J. ; Li, X.-N. ; Crawford, J. R. ; Levy, M. L. ; Kool, M. ; Pfister, S. ; Taylor, M. D. ; Wechsler-Reya, R. J.
Pearson, A. D. J. ; Herold, R. ; Rousseau, R. ; Copland, C. ; Bradley-Garelik, B. ; Binner, D. ; Capdeville, R. ; Caron, H. ; Carleer, J. ; Chesler, L. ; Geoerger, B. ; Kearns, P. ; Marshall, L. V. ; Pfister, S. ; Schleiermacher, G. ; Skolnik, J. ; Spadoni, C. ; Sterba, J. ; van den Berg, H. ; Uttenreuther-Fischer, M. ; Witt, O. ; Norga, K. ; Vassal, G. ; ACCELERATE, M. o. W. G. 1. o. t. P. P. o. ; Georger, B. ; Iannone, R. ; Jakacki, R. ; Russo, M.
Oh, S. ; Flynn, R. A. ; Floor, S. N. ; Purzner, J. ; Martin, L. ; Do, B. T. ; Schubert, S. ; Vaka, D. ; Morrissy, S. ; Li, Y. ; Kool, M. ; Hovestadt, V. ; Jones, D. ; Northcott, P. A. ; Risch, T. ; Warnatz, H.-J. ; Yaspo, M.-L. ; Adams, C. M. ; Leib, R. D. ; Breese, M. ; Marra, M. A. ; Malkin, D. ; Lichter, P. ; Doudna, J. A. ; Pfister, S. ; Taylor, M. D. ; Chang, H. Y. ; Cho, Y.-J.
Lin, C. Y. ; Erkek, S. ; Tong, Y. ; Yin, L. ; Federation, A. J. ; Zapatka, M. ; Haldipur, P. ; Kawauchi, D. ; Risch, T. ; Warnatz, H.-J. ; Worst, B. ; Ju, B. ; Orr, B. A. ; Zeid, R. ; Polaski, D. R. ; Segura-Wang, M. ; Waszak, S. M. ; Jones, D. ; Kool, M. ; Hovestadt, V. ; Buchhalter, I. ; Sieber, L. ; Johann, P. ; Chavez, L. ; Gröschel, S. ; Ryzhova, M. ; Korshunov, A. ; Chen, W. ; Chizhikov, V. V. ; Millen, K. J. ; Amstislavskiy, V. ; Lehrach, H. ; Yaspo, M.-L. ; Eils, R. ; Lichter, P. ; Korbel, J. O. ; Pfister, S. ; Bradner, J. E. ; Northcott, P. A.
Li, P. ; Lee, E. H. ; Du, F. ; Gordon, R. E. ; Yuelling, L. W. ; Liu, Y. ; Ng, J. M. Y. ; Zhang, H. ; Wu, J. ; Korshunov, A. ; Pfister, S. ; Curran, T. ; Yang, Z.-J.
Korshunov, A. ; Capper, D. ; Reuss, D. ; Schrimpf, D. ; Ryzhova, M. ; Hovestadt, V. ; Sturm, D. ; Meyer, J. ; Jones, C. ; Zheludkova, O. ; Kumirova, E. ; Golanov, A. ; Kool, M. ; Schüller, U. ; Mittelbronn, M. ; Hasselblatt, M. ; Schittenhelm, J. ; Reifenberger, G. ; Herold-Mende, C. ; Lichter, P. ; von Deimling, A. ; Pfister, S. ; Jones, D.
Keil, M. ; Sonner, J. ; Lanz, T. ; Oezen, I. ; Bunse, T. ; Bittner, S. ; Meyer, H. V. ; Meuth, S. G. ; Wick, W. ; Platten, M.
Heim, S. ; Sill, M. ; Jones, D. ; Vasiljevic, A. ; Jouvet, A. ; Fèvre-Montange, M. ; Wesseling, P. ; Beschorner, R. ; Mittelbronn, M. ; Kohlhof, P. ; Hovestadt, V. ; Johann, P. ; Kool, M. ; Pajtler, K. ; Korshunov, A. ; Ruland, V. ; Sperveslage, J. ; Thomas, C. ; Witt, H. ; von Deimling, A. ; Paulus, W. ; Pfister, S. ; Capper, D. ; Hasselblatt, M.
Hasselblatt, M. ; Thomas, C. ; Hovestadt, V. ; Schrimpf, D. ; Johann, P. ; Bens, S. ; Oyen, F. ; Peetz-Dienhart, S. ; Crede, Y. ; Wefers, A. ; Vogel, H. ; Riemenschneider, M. J. ; Antonelli, M. ; Giangaspero, F. ; Bernardo, M. C. ; Giannini, C. ; Ud Din, N. ; Perry, A. ; Keyvani, K. ; van Landeghem, F. ; Sumerauer, D. ; Hauser, P. ; Capper, D. ; Korshunov, A. ; Jones, D. ; Pfister, S. ; Schneppenheim, R. ; Siebert, R. ; Frühwald, M. C. ; Kool, M.
Gessi, M. ; Capper, D. ; Sahm, F. ; Huang, K. ; von Deimling, A. ; Tippelt, S. ; Fleischhack, G. ; Scherbaum, D. ; Alfer, J. ; Juhnke, B.-O. ; von Hoff, K. ; Rutkowski, S. ; Warmuth-Metz, M. ; Chavez, L. ; Pfister, S. ; Pietsch, T. ; Jones, D. ; Sturm, D.
Ernst, A. ; Jones, D. ; Maass, K. ; Rode, A. ; Deeg, K. ; Jebaraj, B. M. C. ; Korshunov, A. ; Hovestadt, V. ; Tainsky, M. A. ; Pajtler, K. ; Bender, S. ; Brabetz, S. ; Gröbner, S. ; Kool, M. ; Devens, F. ; Edelmann, J. ; Zhang, C. ; Castelo-Branco, P. ; Tabori, U. ; Malkin, D. ; Rippe, K. ; Stilgenbauer, S. ; Pfister, S. ; Zapatka, M. ; Lichter, P.
D'Asti, E. ; Huang, A. ; Kool, M. ; Meehan, B. ; Chan, J. A. ; Jabado, N. ; Korshunov, A. ; Pfister, S. ; Rak, J.
Chagtai, T. ; Zill, C. ; Dainese, L. ; Wegert, J. ; Savola, S. ; Popov, S. ; Mifsud, W. ; Vujanić, G. ; Sebire, N. ; Le Bouc, Y. ; Ambros, P. F. ; Kager, L. ; O'Sullivan, M. J. ; Blaise, A. ; Bergeron, C. ; Mengelbier, L. H. ; Gisselsson, D. ; Kool, M. ; Tytgat, G. A. M. ; van den Heuvel-Eibrink, M. M. ; Graf, N. ; van Tinteren, H. ; Coulomb, A. ; Gessler, M. ; Williams, R. D. ; Pritchard-Jones, K.
Carceller, F. ; Fowkes, L. A. ; Khabra, K. ; Moreno, L. ; Saran, F. ; Burford, A. ; Mackay, A. ; Jones, D. ; Hovestadt, V. ; Marshall, L. V. ; Vaidya, S. ; Mandeville, H. ; Jerome, N. ; Bridges, L. R. ; Laxton, R. ; Al-Sarraj, S. ; Pfister, S. ; Leach, M. O. ; Pearson, A. D. J. ; Jones, C. ; Koh, D.-M. ; Zacharoulis, S.
Woehrer, A. ; Lau, C. C. ; Prayer, D. ; Bauchet, L. ; Rosenfeld, M. ; Capper, D. ; Fisher, P. G. ; Kool, M. ; Müller, M. ; Kros, J. M. ; Kruchko, C. ; Wiemels, J. ; Wrensch, M. ; Danysh, H. E. ; Zouaoui, S. ; Heck, J. E. ; Johnson, K. J. ; Qi, X. ; O'Neill, B. P. ; Afzal, S. ; Scheurer, M. E. ; Bainbridge, M. N. ; Nousome, D. ; Bahassi, E. M. ; Hainfellner, J. A. ; Barnholtz-Sloan, J. S. ; Consortium, B. T. E.
Wegert, J. ; Ishaque, N. ; Vardapour, R. ; Geörg, C. ; Gu, Z. ; Bieg, M. ; Ziegler, B. ; Bausenwein, S. ; Nourkami, N. ; Ludwig, N. ; Keller, A. ; Grimm, C. ; Kneitz, S. ; Williams, R. D. ; Chagtai, T. ; Pritchard-Jones, K. ; van Sluis, P. ; Volckmann, R. ; Koster, J. ; Versteeg, R. ; Acha, T. ; O'Sullivan, M. J. ; Bode, P. K. ; Niggli, F. ; Tytgat, G. A. ; van Tinteren, H. ; van den Heuvel-Eibrink, M. M. ; Meese, E. ; Vokuhl, C. ; Leuschner, I. ; Graf, N. ; Eils, R. ; Pfister, S. ; Kool, M. ; Gessler, M.
Wang, X. ; Dubuc, A. M. ; Ramaswamy, V. ; Mack, S. ; Gendoo, D. M. A. ; Remke, M. ; Wu, X. ; Garzia, L. ; Luu, B. ; Cavalli, F. ; Peacock, J. ; López, B. ; Skowron, P. ; Zagzag, D. ; Lyden, D. ; Hoffman, C. ; Cho, Y.-J. ; Eberhart, C. ; MacDonald, T. ; Li, X.-N. ; Van Meter, T. ; Northcott, P. A. ; Haibe-Kains, B. ; Hawkins, C. ; Rutka, J. T. ; Bouffet, E. ; Pfister, S. ; Korshunov, A. ; Taylor, M. D.
Sandén, E. ; Dyberg, C. ; Krona, C. ; Visse, E. ; Carén, H. ; Northcott, P. A. ; Kool, M. ; Ståhl, N. ; Persson, A. ; Englund, E. ; Johnsen, J. I. ; Siesjö, P. ; Darabi, A.
Reuss, D. E. ; Kratz, A. ; Sahm, F. ; Capper, D. ; Schrimpf, D. ; Koelsche, C. ; Hovestadt, V. ; Bewerunge-Hudler, M. ; Jones, D. ; Schittenhelm, J. ; Mittelbronn, M. ; Rushing, E. ; Simon, M. ; Westphal, M. ; Unterberg, A. ; Platten, M. ; Paulus, W. ; Reifenberger, G. ; Tonn, J.-C. ; Aldape, K. ; Pfister, S. ; Korshunov, A. ; Weller, M. ; Herold-Mende, C. ; Wick, W. ; Brandner, S. ; von Deimling, A.
Merino, D. M. ; Shlien, A. ; Villani, A. ; Pienkowska, M. ; Mack, S. ; Ramaswamy, V. ; Shih, D. ; Tatevossian, R. ; Novokmet, A. ; Choufani, S. ; Dvir, R. ; Ben-Arush, M. ; Harris, B. T. ; Hwang, E. I. ; Lulla, R. ; Pfister, S. ; Achatz, M. I. ; Jabado, N. ; Finlay, J. L. ; Weksberg, R. ; Bouffet, E. ; Hawkins, C. ; Taylor, M. D. ; Tabori, U. ; Ellison, D. W. ; Gilbertson, R. J. ; Malkin, D.
Mack, S. C. ; Agnihotri, S. ; Bertrand, K. C. ; Wang, X. ; Shih, D. J. ; Witt, H. ; Hill, N. ; Zayne, K. ; Barszczyk, M. ; Ramaswamy, V. ; Remke, M. ; Thompson, Y. ; Ryzhova, M. ; Massimi, L. ; Grajkowska, W. ; Lach, B. ; Gupta, N. ; Weiss, W. A. ; Guha, A. ; Hawkins, C. ; Croul, S. ; Rutka, J. T. ; Pfister, S. ; Korshunov, A. ; Pekmezci, M. ; Tihan, T. ; Philips, J. J. ; Jabado, N. ; Zadeh, G. ; Taylor, M. D.
Korshunov, A. ; Ryzhova, M. ; Hovestadt, V. ; Bender, S. ; Sturm, D. ; Capper, D. ; Meyer, J. ; Schrimpf, D. ; Kool, M. ; Northcott, P. A. ; Zheludkova, O. ; Milde, T. ; Witt, O. ; Kulozik, A. E. ; Reifenberger, G. ; Jabado, N. ; Perry, A. ; Lichter, P. ; von Deimling, A. ; Pfister, S. ; Jones, D.
Koelsche, C. ; Hovestadt, V. ; Jones, D. ; Capper, D. ; Sturm, D. ; Sahm, F. ; Schrimpf, D. ; Adeberg, S. ; Böhmer, K. ; Hagenlocher, C. ; Mechtersheimer, G. ; Kohlhof, P. ; Mühleisen, H. ; Beschorner, R. ; Hartmann, C. ; Braczynski, A. K. ; Mittelbronn, M. ; Buslei, R. ; Becker, A. ; Grote, A. ; Urbach, H. ; Staszewski, O. ; Prinz, M. ; Hewer, E. ; Pfister, S. ; von Deimling, A. ; Reuss, D. E.
Giachino, C. ; Boulay, J.-L. ; Ivanek, R. ; Alvarado, A. ; Tostado, C. ; Lugert, S. ; Tchorz, J. ; Coban, M. ; Mariani, L. ; Bettler, B. ; Lathia, J. ; Frank, S. ; Pfister, S. ; Kool, M. ; Taylor, V.
Geisenberger, C. ; Mock, A. ; Warta, R. ; Rapp, C. ; Schwager, C. ; Korshunov, A. ; Nied, A.-K. ; Capper, D. ; Brors, B. ; Jungk, C. ; Jones, D. ; Collins, V. P. ; Ichimura, K. ; Bäcklund, L. M. ; Schnabel, E. ; Mittelbron, M. ; Lahrmann, B. ; Zheng, S. ; Verhaak, R. G. W. ; Grabe, N. ; Pfister, S. ; Hartmann, C. ; von Deimling, A. ; Debus, J. ; Unterberg, A. ; Abdollahi, A. ; Herold-Mende, C.
Gallo, M. ; Coutinho, F. J. ; Vanner, R. J. ; Gayden, T. ; Mack, S. C. ; Murison, A. ; Remke, M. ; Li, R. ; Takayama, N. ; Desai, K. ; Lee, L. ; Lan, X. ; Park, N. I. ; Barsyte-Lovejoy, D. ; Smil, D. ; Sturm, D. ; Kushida, M. M. ; Head, R. ; Cusimano, M. D. ; Bernstein, M. ; Clarke, I. D. ; Dick, J. E. ; Pfister, S. ; Rich, J. N. ; Arrowsmith, C. H. ; Taylor, M. D. ; Jabado, N. ; Bazett-Jones, D. P. ; Lupien, M. ; Dirks, P. B.
Fontebasso, A. M. ; Shirinian, M. ; Khuong-Quang, D.-A. ; Bechet, D. ; Gayden, T. ; Kool, M. ; De Jay, N. ; Jacob, K. ; Gerges, N. ; Hutter, B. ; Şeker-Cin, H. ; Witt, H. ; Montpetit, A. ; Brunet, S. ; Lepage, P. ; Bourret, G. ; Klekner, A. ; Bognár, L. ; Hauser, P. ; Garami, M. ; Farmer, J.-P. ; Montes, J.-L. ; Atkinson, J. ; Lambert, S. ; Kwan, T. ; Korshunov, A. ; Tabori, U. ; Collins, V. P. ; Albrecht, S. ; Faury, D. ; Pfister, S. ; Paulus, W. ; Hasselblatt, M. ; Jones, D. ; Jabado, N.
Faria, C. C. ; Golbourn, B. J. ; Dubuc, A. M. ; Remke, M. ; Diaz, R. J. ; Agnihotri, S. ; Luck, A. ; Sabha, N. ; Olsen, S. ; Wu, X. ; Garzia, L. ; Ramaswamy, V. ; Mack, S. C. ; Wang, X. ; Leadley, M. ; Reynaud, D. ; Ermini, L. ; Post, M. ; Northcott, P. A. ; Pfister, S. ; Croul, S. E. ; Kool, M. ; Korshunov, A. ; Smith, C. A. ; Taylor, M. D. ; Rutka, J. T.
Betts, M. J. ; Lu, Q. ; Jiang, Y. ; Drusko, A. ; Wichmann, O. ; Utz, M. ; Valtierra-Gutiérrez, I. A. ; Schlesner, M. ; Jaeger, N. ; Jones, D. ; Pfister, S. ; Lichter, P. ; Eils, R. ; Siebert, R. ; Bork, P. ; Apic, G. ; Gavin, A.-C. ; Russell, R. B.
An, J. ; González-Avalos, E. ; Chawla, A. ; Jeong, M. ; López-Moyado, I. F. ; Li, W. ; Goodell, M. A. ; Chavez, L. ; Ko, M. ; Rao, A.
Vanner, R. J. ; Remke, M. ; Gallo, M. ; Selvadurai, H. J. ; Coutinho, F. ; Lee, L. ; Kushida, M. ; Head, R. ; Morrissy, S. ; Zhu, X. ; Aviv, T. ; Voisin, V. ; Clarke, I. D. ; Li, Y. ; Mungall, A. J. ; Moore, R. A. ; Ma, Y. ; Jones, S. J. M. ; Marra, M. A. ; Malkin, D. ; Northcott, P. A. ; Kool, M. ; Pfister, S. ; Bader, G. ; Hochedlinger, K. ; Korshunov, A. ; Taylor, M. D. ; Dirks, P. B.
Wiestler, B. P. O. ; Capper, D. ; Sill, M. ; Jones, D. ; Hovestadt, V. ; Sturm, D. ; Koelsche, C. ; Bertoni, A. ; Schweizer, L. ; Korshunov, A. ; Weiß, E. K. ; Schliesser, M. ; Radbruch, A. ; Herold-Mende, C. ; Roth, P. ; Unterberg, A. ; Hartmann, C. ; Pietsch, T. ; Reifenberger, G. ; Lichter, P. ; Radlwimmer, B. ; Platten, M. ; Pfister, S. ; von Deimling, A. ; Weller, M. ; Wick, W.
Sturm, D. ; Bender, S. ; Jones, D. ; Lichter, P. ; Grill, J. ; Becher, O. ; Hawkins, C. ; Majewski, J. ; Jones, C. ; Costello, J. F. ; Iavarone, A. ; Aldape, K. ; Brennan, C. W. ; Jabado, N. ; Pfister, S.
Sahm, F. ; Reuss, D. ; Koelsche, C. ; Capper, D. ; Schittenhelm, J. ; Heim, S. ; Jones, D. ; Pfister, S. ; Herold-Mende, C. ; Wick, W. ; Mueller, W. ; Hartmann, C. ; Paulus, W. ; von Deimling, A.
Korshunov, A. ; Sturm, D. ; Ryzhova, M. ; Hovestadt, V. ; Gessi, M. ; Jones, D. ; Remke, M. ; Northcott, P. ; Perry, A. ; Picard, D. ; Rosenblum, M. ; Antonelli, M. ; Aronica, E. ; Schüller, U. ; Hasselblatt, M. ; Woehrer, A. ; Zheludkova, O. ; Kumirova, E. ; Puget, S. ; Taylor, M. D. ; Giangaspero, F. ; Peter Collins, V. ; von Deimling, A. ; Lichter, P. ; Huang, A. ; Pietsch, T. ; Pfister, S. ; Kool, M.
Karajannis, M. A. ; Legault, G. ; Fisher, M. J. ; Milla, S. S. ; Cohen, K. J. ; Wisoff, J. H. ; Harter, D. H. ; Goldberg, J. D. ; Hochman, T. ; Merkelson, A. ; Bloom, M. C. ; Sievert, A. J. ; Resnick, A. C. ; Dhall, G. ; Jones, D. ; Korshunov, A. ; Pfister, S. ; Eberhart, C. G. ; Zagzag, D. ; Allen, J. C.
Ramaswamy, V. ; Remke, M. ; Shih, D. ; Wang, X. ; Northcott, P. A. ; Faria, C. C. ; Raybaud, C. ; Tabori, U. ; Hawkins, C. ; Rutka, J. ; Taylor, M. D. ; Bouffet, E.
Rack, P. G. ; Ni, J. ; Payumo, A. Y. ; Nguyen, V. ; Crapster, J. A. ; Hovestadt, V. ; Kool, M. ; Jones, D. ; Mich, J. K. ; Firestone, A. J. ; Pfister, S. ; Cho, Y.-J. ; Chen, J. K.
Hovestadt, V. ; Jones, D. ; Picelli, S. ; Wang, W. ; Kool, M. ; Northcott, P. A. ; Sultan, M. ; Stachurski, K. ; Ryzhova, M. ; Warnatz, H.-J. ; Ralser, M. ; Brun, S. ; Bunt, J. ; Jäger, N. ; Kleinheinz, K. ; Erkek, S. ; Weber, U. ; Bartholomae, C. C. ; von Kalle, C. ; Lawerenz, C. ; Eils, J. ; Koster, J. ; Versteeg, R. ; Milde, T. ; Witt, O. ; Schmidt, S. ; Wolf, S. ; Pietsch, T. ; Rutkowski, S. ; Scheurlen, W. ; Taylor, M. D. ; Brors, B. ; Felsberg, J. ; Reifenberger, G. ; Borkhardt, A. ; Lehrach, H. ; Wechsler-Reya, R. J. ; Eils, R. ; Yaspo, M.-L. ; Landgraf, P. ; Korshunov, A. ; Zapatka, M. ; Radlwimmer, B. ; Pfister, S. ; Lichter, P.
He, X. ; Zhang, L. ; Chen, Y. ; Remke, M. ; Shih, D. ; Lu, F. ; Wang, H. ; Deng, Y. ; Yu, Y. ; Xia, Y. ; Wu, X. ; Ramaswamy, V. ; Hu, T. ; Wang, F. ; Zhou, W. ; Burns, D. K. ; Kim, S. H. ; Kool, M. ; Pfister, S. ; Weinstein, L. S. ; Pomeroy, S. L. ; Gilbertson, R. J. ; Rubin, J. B. ; Hou, Y. ; Wechsler-Reya, R. ; Taylor, M. D. ; Lu, Q. R.
Pöschl, J. ; Stark, S. ; Neumann, P. ; Gröbner, S. ; Kawauchi, D. ; Jones, D. ; Northcott, P. A. ; Lichter, P. ; Pfister, S. ; Kool, M. ; Schüller, U.
Pietsch, T. ; Schmidt, R. ; Remke, M. ; Korshunov, A. ; Hovestadt, V. ; Jones, D. ; Felsberg, J. ; Kaulich, K. ; Goschzik, T. ; Kool, M. ; Northcott, P. A. ; von Hoff, K. ; von Bueren, A. O. ; Friedrich, C. ; Mynarek, M. ; Skladny, H. ; Fleischhack, G. ; Taylor, M. D. ; Cremer, F. ; Lichter, P. ; Faldum, A. ; Reifenberger, G. ; Rutkowski, S. ; Pfister, S.
Get in touch with us


