Mechanisms of Genome Control

Dr. Angelika Feldmann
Group Leader
We investigate how genes are regulated in space and time and how these mechanisms are misdirected in cancer.
Image: [Translate to English:] Image: Visualization of 3D genome structure as Hi-C (top), Capture-C (middle) and cartoon (bottom),
![[Translate to English:]](/fileadmin/_processed_/9/b/csm_Research_Profile_picture_1e17f36da6.webp)
Image: [Translate to English:] Image: Visualization of 3D genome structure as Hi-C (top), Capture-C (middle) and cartoon (bottom),
Our Research
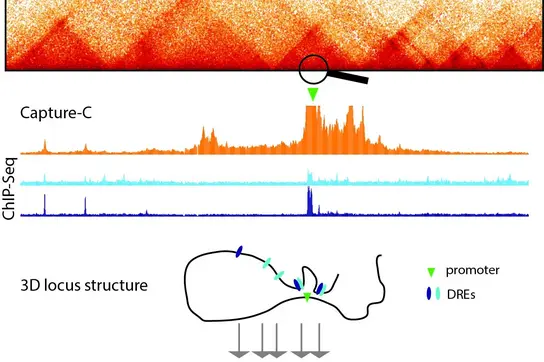
The Feldmann lab studies how genes are regulated in time and space, and how these programs become corrupted in cancers. One of the most fascinating processes in biology is the development of a multicellular organism with >200 different cell types from just one single cell. This means that most of these cell types have exactly the same genetic content despite being highly diverse in their appearance and function. Such diversity is only possible if genes are precisely regulated in space and time. To achieve this, gene promoters must integrate inputs from distal gene regulatory elements (DREs), such as enhancers. Despite extensive studies, we still lack a general understanding about the mechanisms behind DRE-driven gene activation.
Often located hundreds of kilobases away, DREs are thought to come into close physical proximity to their target genes, which involves folding of the DNA and is regulated by a number of proteins (Schoenfelder et al., 2019, Feldmann et al., 2020, Rhodes et al., 2020). Such physical interactions correlate with gene activity and are frequently altered in cancer, suggesting that they may be required for transcriptional alterations observed in normal development and carcinogenesis. Recent data, however, reveal that physical contacts between DREs and gene promoters can occur independently of gene activity (Ghavi-Helm et al., 2014; Benabdallah et al., 2019, Feldmann et al., 2020, Mahara et al., 2024), indicating that other mechanisms underlie their function in gene activation.
Our mission is to understand the precise mechanisms by which DREs communicate with gene promoters during activation and maintenance of transcription, with a specific focus on the role of the 3D genome structure in this process. We combine state-of-the-art genomic, proteomic and computational biology approaches with high-throughput genetic engineering to dissect gene regulatory mechanisms in health and disease. We use embryonic stem cell differentiation and cancer cell lines, which are ideally suitable for genetic manipulation, as models for gene activation.
Selected Publications
S Mahara, S Prüssing, V Smialkovska, S Krall, S Holliman, B Blum, V Dachtler, H Borgers, E Sollier, C Plass and A Feldmann
Petersen Frido*, Westermann Simon*, Smialkovska Valeriia*, Mathony Jan, Feldmann Angelika#, Niopek Dominik#
Feldmann Angelika#, Dimitrova, Emilia, Kenney Alexander, Lastuvkova Anna, Klose Robert J#
Rhodes, J. D. P.*, Feldmann, A.*, Hernandez-Rodriguez*, B., Diaz*, N., Brown, J. M., Fursova, N. A., Blackledge, N. P., Prathapan, P., Dobrinic, P., Huseyin, M. K., Szczurek, A., Kruse, K., Nasmyth, K. A., Buckle, V. J., Vaquerizas, J. M.#, Klose, R. J#
Scientific and other news
2025
December: Second graduation from our lab: Congratulations, Nick, for defending your Master Thesis!
November: Congratulations to our fantastic postdoc Ann-Kristin for receiving the publication award from Deutsche Gesellschaft für Andrologie e.V.
November: Our second review is out: Engineering the Link: From Genome Interaction Maps to Functional Insight. We discuss how to manipulate enhancer-promoter interactions and why we would even need to do this Really cool collaboration with Dominik Niopek and his lab. Congratulations to the amazing first authors Frido, Simon and Valeriia. Looking forward to our joint projects!
October: Cassy joins the lab for a master rotation!
June: Welcome, Frido, who joins the lab for a master internship and Julia, our short-term technician.
May: Welcome to our Master students, Nick and Gaia.
January: The first review from the lab is out, featuring our view on the role of enhancers, 3D genome structure, and timing in both normal development and cancer: Enhancer-dependent gene regulation in space, time, and malignancies. Great work from the joint first authors, Vici and Belinda.
2024
November: Congratulations to our new postdoc Ann-Kristin for receiving the Dr. Rurainski Fellowship
October: The first paper of the lab is out!!!! We find that the dynamics of gene activation in development is characterized by a distinct dynamics of genome folding and entails transient interactions. Congratulations to our first author Sylvia, all the present and past members of the lab, as well as our collaborators from Christoph Plass' lab for this truly collaborative effort! We celebrate with a glass of champagne!
October: Gaia joins the lab for a master internship.
October, 1st: Our lab is 3 years old! We celebrate with a potluck lunch, Escape Room challenge and a dinner
June: Ali, our Erasmus Summer student joins the lab.
June: Official start of our ERC project TimedEnhancer. We are looking forward to make important contributions to the field and are grateful for the trust!
April: Welcome to Belinda, our 3rd PhD student!
March: Double-celebration: Congratulations to our amazing PhD students Vici for receiving the DKFZ PhD Fellowship and Valeriia for receiving the prize for best participation and presentation at the Paris Course of Epigenetics!!!
February: We are saying goodbye to Sylvia on her new adventure in Australia after a productive postdoc! Your work was fundamental for the beginnings of our lab, thank you!
February: Welcome to our newest PhD student Vici!
2023
December: The very first paper with our lab as coauthors is out. Happy to have contributed to this beautiful study by Maja Funk from Michael Boutros' lab!
December: We recieve the ERC Starting grant!
September: Welcome back to Valeriia, who is starting her PhD with us!
April: We celebrate Valeriia's Master thesis the first thesis defense of the lab
February: We say goodbye to Chloe and welcome our first Erasmus student Sam
2022
July: We welcome our first Master student, Valeriia, and our new technician, Sonja, with the first joint lab retreat together with Pei-Chi Wei's and Lena Kutscher's labs.
June: Goodbye to the very first member of the lab, Helena. It was apleasure setting up the lab with you!
March: Welcome to the first postdoc of the lab, Sylvia!
February: Welcome to Chloe!
2021
October: Our lab opens its doors with its first member, Helena!
2025
Engineering the Link: From Genome Interaction Maps to Functional Insight. Adv Biol (Weinh). Petersen, F.*, Westermann, S.*, Smialkovska, V.*, Mathony, J., Feldmann, A.#, Niopek, D.#
Enhancer-dependent gene regulation in space, time, and malignancies. Int J Cancer. Blum, B.*, V. Dachtler*, and A. Feldmann. 2025.
2024
Transient promoter interactions modulate developmental gene activation. Mol Cell. 84:4486-4502 e4487. Mahara, S., S. Prussing, V. Smialkovska, S. Krall, S. Holliman, B. Blum, V. Dachtler, H. Borgers, E. Sollier, C. Plass, and A. Feldmann. 2024
Anti-phase clustering of regulatory factors shapes gene bursting. bioRxiv. Li, B., Y.Y. Wong, N. Flores-Rodriguez, T. Davidson, M.S. Graus, V. Smialkovska, H. Ohishi, A. Feldmann, H. Ochiai, and M. Francois. 2024
2023
Aged intestinal stem cells propagate cell-intrinsic sources of inflammaging in mice. Dev Cell. 58:2914-2929 e2917. Funk, M.C., J.G. Gleixner, F. Heigwer, D. Vonficht, E. Valentini, Z. Aydin, E. Tonin, S. Del Prete, S. Mahara, Y. Throm, J. Hetzer, D. Heide, O. Stegle, D.T. Odom, A. Feldmann, S. Haas, M. Heikenwalder, and M. Boutros. 2023.
Changes in PRC1 activity during interphase modulate lineage transition in pluripotent cells. Nat Commun. 14:180. Asenjo, H.G., M. Alcazar-Fabra, M. Espinosa-Martinez, L. Lopez-Onieva, A. Gallardo, E. Dimitrova, A. Feldmann, T. Pachano, J. Martorell-Marugan, P. Carmona-Saez, A. Sanchez-Pozo, A. Rada-Iglesias, R.J. Klose, and D. Landeira. 2023.
2022
Distinct roles for CKM-Mediator in controlling Polycomb-dependent chromosomal interactions and priming genes for induction. Nat Struct Mol Biol. 29:1000-1010. Dimitrova, E., A. Feldmann, R.H. van der Weide, K.D. Flach, A. Lastuvkova, E. de Wit, and R.J. Klose. 2022.
2020
A genome-scale map of DNA methylation turnover identifies site-specific dependencies of DNMT and TET activity. Nat Commun. 11:2680. Ginno, P.A., D. Gaidatzis, A. Feldmann, L. Hoerner, D. Imanci, L. Burger, F. Zilbermann, A. Peters, F. Edenhofer, S.A. Smallwood, A.R. Krebs, and D. Schubeler. 2020.
CDK-Mediator and FBXL19 prime developmental genes for activation by promoting atypical regulatory interactions. Nucleic Acids Res. 48:2942-2955. Feldmann, A., E. Dimitrova, A. Kenney, A. Lastuvkova, and R.J. Klose. 2020.
Cohesin Disrupts Polycomb-Dependent Chromosome Interactions in Embryonic Stem Cells. Cell Rep. 30:820-835 e810. Rhodes, J.D.P., A. Feldmann, B. Hernandez-Rodriguez, N. Diaz, J.M. Brown, N.A. Fursova, N.P. Blackledge, P. Prathapan, P. Dobrinic, M.K. Huseyin, A. Szczurek, K. Kruse, K.A. Nasmyth, V.J. Buckle, J.M. Vaquerizas, and R.J. Klose. 2020.
PRC1 Catalytic Activity Is Central to Polycomb System Function. Mol Cell. 77:857-874 e859. Blackledge, N.P., N.A. Fursova, J.R. Kelley, M.K. Huseyin, A. Feldmann, and R.J. Klose. 2020.
2018
FBXL19 recruits CDK-Mediator to CpG islands of developmental genes priming them for activation during lineage commitment. Elife. 7. Dimitrova, E., T. Kondo, A. Feldmann, M. Nakayama, Y. Koseki, R. Konietzny, B.M. Kessler, H. Koseki, and R.J. Klose. 2018.
2017
The SET1 Complex Selects Actively Transcribed Target Genes via Multivalent Interaction with CpG Island Chromatin. Cell Rep. 20:2313-2327. Brown, D.A., V. Di Cerbo, A. Feldmann, J. Ahn, S. Ito, N.P. Blackledge, M. Nakayama, M. McClellan, E. Dimitrova, A.H. Turberfield, H.K. Long, H.W. King, S. Kriaucionis, L. Schermelleh, T.G. Kutateladze, H. Koseki, and R.J. Klose. 2017.
2014
Acetylation of histone H3 at lysine 64 regulates nucleosome dynamics and facilitates transcription. Elife. 3:e01632. Di Cerbo, V., F. Mohn, D.P. Ryan, E. Montellier, S. Kacem, P. Tropberger, E. Kallis, M. Holzner, L. Hoerner, A. Feldmann, F.M. Richter, A.J. Bannister, G. Mittler, J. Michaelis, S. Khochbin, R. Feil, D. Schuebeler, T. Owen-Hughes, S. Daujat, and R. Schneider. 2014.
2013
Transcription factor occupancy can mediate active turnover of DNA methylation at regulatory regions. PLoS Genet. 9:e1003994. Feldmann, A., R. Ivanek, R. Murr, D. Gaidatzis, L. Burger, and D. Schubeler. 2013.
2024
Presentation: Akademische Mittagspause. Kommunikation auf Distanz: Wie das Genom die Entwicklung reguliert
2023
Press release: ERC Starting Grant for Angelika Feldmann
Interview #EmbraceEquity
2021
Interview: 100 Women in Science at Oxford University
2020
Presentation: FragileNucleosome. Cohesin disrupts polycomb-dependent chromosome interactions in ESCs.
Lab Resources
Our lab seeks to understand how genome structure and function are connected. In order to decipher causal relationships we will develop unique methods that will allow us to manipulate genome structure in healthy and cancerous cells with the future goal to transform personalised cancer treatment. An important focus of the lab is to mentor the next generation of scientists. We will create a collaborative, respectful atmosphere, conductive of independent learning and thinking. We want to be known in the community as fair, rigorous and innovative.
Feldmann lab expectations:
Below we've listed some of the things that you can expect from Angelika and what she is expecting from her lab members. Having an understanding of our mutual expectations is key to the success of a mentoring relationship. This is meant to be a dynamic document that we will continue to revise collaboratively.
Please note that these expectations were heavily influenced/inspired by those posted by Elena Kramer.
Your expectations of Angelika:
- I am very committed to mentoring and will be trying to provide valuable advice for your particular situation. You can expect me to put much energy and effort into developing you as a scientist. I will also particularly strive to help you with your career development, or guide you towards somebody who can help you, whatever career you would like to pursue. In fact, you can expect me to regularly check in with you regarding your personal career goals. However, no mentor can know everything, so I encourage every lab member to seek out additional mentors.
- You can expect to meet with me regularly for at least 30 minutes a week. I think it's important that you have regular one-on-one time with me. You can expect me to be on time and let you know if I'm running late. I have an open-door policy, so drop-ins and questions or discussions are strongly encouraged.
- You can also expect me to be hands-on with both experimental and computational issues. I will show you some of the new techniques directly. This is part of the advantage of being in a new group.
- You can expect me to keep a close eye on you whenever you are doing something for the first time. This is not a critique on your technique, but a reflection of me having to ensure that methods are set-up properly in our new group.
- You can expect me to be particularly picky on some of the experimental techniques (such as tissue culture). This is to make sure everyone does the key techniques in the same way and uses the same protocol. Changes in the protocol will have to be discussed with everyone and brought up in the lab meetings. This will ensure high reproducibility and a simpler trouble-shooting within the lab.
- You can expect me to ask for your honest feedback about the lab, your experience, and your perception of me as a mentor. I can't improve something if you don't tell me about it (and I really do want to improve things).
- You can expect me to give you my honest feedback about your progress and work style. I am here to support you in every way possible and giving you feedback is one of the most important ways to promote your success.
- You can expect me to help you set reasonable deadlines for making progress and set clear expectations regarding what that progress should be - what do we want and when do we want it. You can then expect me to keep a rough eye on your timeline. Ultimately, you are responsible for making sure you hit the deadlines, but I will help you plan things out so that there are no surprises. Good planning will help you to work more efficiently and achieve a better work-life balance. I will closely monitor your progress and actively help you improve it if it is not ideal.
- You can expect me to coach you to become independent while maintaining high standards. This means that while you will initially have a close supervision it will become progressively more hands-off as you settle into the lab. The timeline depends on your progress, accuracy and comfort with working independently. If I notice that your progress stalls with less input from my side you can expect me to supervise you more closely for a while. Postdocs can expect to have largely conceptual input, unless they ask for more detailed advice.
- You can expect me to encourage you to pursue your own curiosity-driven pet and side projects that take up ~20% of your time.
- You can expect me to provide timely feedback on your TAC reports, grants and papers. 1-2 weeks turn-around time is fair unless it is particularly complicated.
- I will try to respond to emails within 24-hours during the work week. If you don't hear from me by then, feel free to send a reminder.
- You can expect me to listen to your concerns and help you solve problems. Some problems need to be strong-armed and when that is the case, I will gladly step in and advocate for you whenever necessary, whether with the department or collaborators. It's my job.
- You can expect me to write letters of recommendation for you, given advance notice. If it's a sort of letter I've never written before or the very first time I've written a letter for you, please give me two weeks lead time because I need to find a 3-4 hour block of time to write you a good letter. After I have written you a letter for something once, a week of lead time is sufficient to ask me to send another (I will tailor a letter I've already written). This being said, I understand that sometimes opportunities crop up unexpectedly. I will do my best under rush circumstances but would not expect it to be the norm.
- You can expect me to help you network. I will write emails to introduce you to people in the field and I will connect you to people at meetings. I will also guide you in knowing what to expect from these interactions (having an 'elevator speech', asking them questions.). These things are more important than you think in getting a position after your degree.
- You can expect me to work very hard with you on your posters and talks. These are your face to the research community and if the audience is giving you their time, you want to make the most of it so I will give you my time to make your talk or poster effective and engaging.
- You can expect me to brainstorm new ideas for projects with you. We can have two modes that we agree upon: A playful mode and a critical mode. Both modes are absolutely necessary. A brainstorming will always first start with a playful mode to collect ideas. However, then I will scrutinize them and invite you to join in. This will make sure we think about all possible solutions and problems.
- You can expect me to help you find funding sources. I will send you all sorts of opportunities and you will decide what to pursue.
- You can expect me to work as hard toward your success as you do. The harder you push, the harder I will push to help.
- You can expect me to ensure that you have a safe and supportive environment in which to work. If you have any concerns about the climate in the lab, I want to know as soon as possible. All information remains confidential unless you choose to release it.
- You can expect me to provide a model of professionalism in academia, and you can look to me for guidance on issues related to conflicts of interest, equity and fairness, ethical research, and mentoring.
- You can expect me to help you find other resources and other mentors when you need additional support beyond my abilities. Your advisor cannot possibly provide all the advice and support you will need in your career, so developing a robust mentoring network early on is very helpful.
Angelika's expectations of you:
- I expect that you will pursue research that has some overlap with my expertise and interests. Otherwise, I will not be in a good position to mentor you. If your interests shift away from the themes of the lab, I expect that you will inform me and we can discuss options, which could include switching to another lab. You should always be open to considering that possibility - again, it's your career, moving to a different lab isn't a problem. Note, I am always open to be convinced about a new interesting idea, but you will need to put effort into your arguments.
- For postdocs fellows, I expect that you will pursue the experiments that we agreed on when you joined the lab as your main project, but I recognize that these plans will evolve and change as the science develops. I am always happy to discuss ideas for new directions, but if my funding is supporting the science, I expect to be closely involved in decisions.
- I expect students and postdocs to keep track of scientific literature relevant to their projects and inform me about any developments that may affect the project. This includes publications that scoop the novelty of your project or suggest a different approach would be much more effective. I consider a regular reading habit to be of critical importance and a key part of your work in the lab. Keep in mind that I will not be able to keep track of all the relevant literature myself and it is key to regularly assess the novelty of your project/approach if you want your work to be publishable and have the best outcomes possible. I expect PhD students and postdocs to spend at least 3 hours per week on reading and thinking about their projects.
- I expect you to make regular short-term (1 week), mid-term (1-3 months) and long-term (1-3 years) plans for your project. This will help you focus and track your progress and will force us to think hard about what we want to achieve. I understand that plans will change and evolve, but good planning is particularly essential to achieve your goals.
- I expect honesty from you. I will not judge you for not knowing something, making a mistake, or for breaking something in the lab. You're here to learn and ask questions. It is unlikely that you will make a mistake that I haven't made myself. At the same time I expect you to learn from your mistakes and to own them. You can expect the same from me.
- I expect you to be resourceful. I don't know all the answers and you may learn more effectively if you figure things out for yourself. So by all means, ask questions, sometimes I will have an answer, and sometimes I will point you to useful paths for answering those questions. At the same time I expect you to not delay asking for my advice if you are not progressing without it.
- I expect you to seek out resources (equipment, chemicals etc.) required for your work. I can help you with that, but only if I know what you need. Nevertheless I expect you to have put some effort into finding the necessary resources yourself.
- I expect that you will respect my time as I respect yours. If we have a meeting scheduled, I expect you to be on time and prepared, or to let me know if you will be late.
- I expect you to be a full participant in this lab community. This means that sometimes you will have to do things for the lab community (buffers, competent cells, setting up a method etc.). That also includes being at lab meetings and joint lab meetings (on time), fully participating in those meetings (by presenting, asking questions), coming to lab social events, taking part in at least some of our outreach events, helping to host visiting speakers, etc. I also expect that you will be supportive of each other -- say hello to your lab mates, help when they ask (if you can), be welcoming to newcomers, etc.
- I expect you to tell me when you can't do one of these things. You are typically juggling just as many things as I am and sometimes have to say no. This is fine and I don't expect you to do everything.
- I expect that, normally, you will be in the lab five days a week. As you know, I'm a morning person, but I don't expect you to work the same hours I do. I'm not going to keep track of when you come in and when you leave, but I do expect to see consistent progress in line with working a full working week, taking into consideration other work-related commitments. When deciding on your working hours, keep in mind that part of the benefit of working in a lab is the ability to learn from other people as part of a scientific community and if you're only here when everyone else is gone, you will be losing out on that.
- If you need to take off it's not a problem, but I expect a personal email from you notifying me about it, which should be sent at least 24 hours in advance if possible.
- I expect that you will usually respond in a timely fashion (24-48 hrs if during the work week) when I email you. I also expect that you will tell me when you will be out of town for extended periods, in case lab issues come up and we are looking for you. Taking holidays is not a problem, but since we are a small lab it is important to notify me on time so that we can plan accordingly who is on-site.
- I expect you to alert lab members and/or appropriate staff if you see something wrong in the lab, whether it be a growth chamber running too hot, a water bath not working, or a computer on the fritz.
- I expect you to complete your assigned lab jobs in a timely manner. The lab only works if everyone pulls their weight. I will know and I will appreciate your efforts. That also includes basic lab cleanliness and common consideration in regard to using shared equipment. If you don't have time to clean up after yourself, you don't have time to do the experiment. If you do notice that certain things aren't getting done, please let me know about it. I don't expect you to monitor or admonish your colleagues. Let me be the enforcer!
- I expect you to let me and/or other lab members know of any complains and concerns in a timely manner. Don't wait until you dwelled on things and they start to really annoy you before you tell someone. In the worst case you will hear the reasons behind the topic of your concern.
- I expect that you will discuss issues of authorship of publications, presentations and talks with me before presenting/submitting. Typically, PIs contribute intellectually and monetarily to student projects and are listed as senior authors. However, if a project did not involve my effort in these ways, I would not expect authorship (and in fact if you do side projects that don't involve me, that's great, but see #3/4 above.)
- I expect you will actively pursue funds to support your research, even if it is in line with ongoing projects in the lab. This is easily one of the most important aspects of your training -- learning how to sell your ideas -- and it takes lots of practice. Your success in obtaining grants will also be one of the main features (beyond your publications) that any future employers will consider, at least in research-related careers. This does not mean that I won't help with paying for your research, but it does mean that I need to see that you are putting in an effort to support yourself.
- I expect you to represent our lab externally and internally within the institute. This includes participating in seminars and conferences (including presentations and questions).
Note: These expectations are specific to me.

Team
- Show profile

Dr. Angelika Feldmann
Group Leader
- Show profile

Belinda Blum
PhD student
- Show profile

Sarah-Victoria Dachtler
PhD student
- Show profile

Dr. Ann-Kristin Dicke
Postdoc
- Show profile

Sonja Prüssing
Technician
- Show profile

Valeriia Smialkovska
PhD student
-
Ezo Özcan
Open Positions
We are continously looking for motivated people to join our lab and we offer positions in various stages.
Postdocs: Please email your application to Dr. Angelika Feldmann and include a cover letter stating your research interest, your CV with a complete list of publications as well as contact details of 3 references. Also check out the International Postdoc Program at the DKFZ.
PhD students: Please apply through the International PhD Program of the DKFZ and also send your application directly to Dr. Angelika Feldmann.
Master students: Please send a cover letter, your CV and transcripts directly to Dr. Angelika Feldmann.
Get in touch with us

