Translational Functional Cancer Genomics
- Functional and Structural Genomics
- NCT

Prof. Dr. Hanno Glimm
Profound patient specific differences in cancer-causing aberrations as well as functional heterogeneity within individual tumors pose major challenges for the development of mechanism-based strategies in clinical cancer care.

Our Research
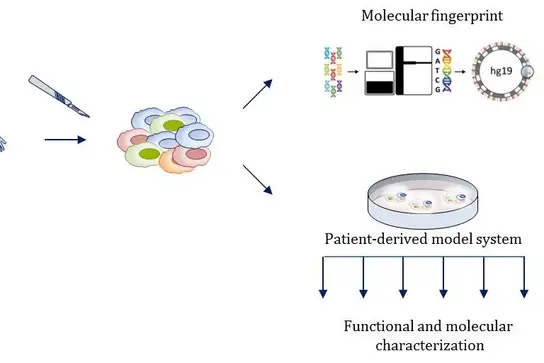
To understand the implications of inter- and intratumoral heterogeneity for personalized treatment approaches scientists and physicians from the Translational Functional Cancer Genomics group study the genomic and functional properties of tumor-cell subpopulations. They use their insights gained from multi-dimensional analysis of individual tumor samples and functional testing of patientderived tumor models to identify and approve novel treatment strategies.
Specific cultures are generated from individual tumor samples of patients that receive comprehensive molecular profiling in the MASTER (Molecularly Aided Stratification for Tumor Eradication) program, to functionally characterize their properties. Using our strong background and expertise in the field of stem cell and molecular cancer biology, we investigate the clonality, heterogeneity and functional characteristics of patient specific primary tumors and their respective model systems, also addressing subpopulation dynamics in benign and malignant stem cell fractions. As tumor-initiating cells very often hijack regulatory mechanisms, which are strictly controlled in benign stem cell systems, we specifically implement high-throughput screening approaches together with state-of-the-art single cell analyses and mouse genetics to delineate critical and deregulated factors leading to tumor initiation, metastasis formation and therapy resistance. Identified mechanistic insights are translated into clinical treatment strategies whenever possible.
The group of Translational Functional Cancer Genomics in Heidelberg joins forces with researchers of the Department of Translational Medical Oncology in Dresden to combine complementary strengths of both NCT partner sides in shared translational research programs.
Team
-

Maximilian Bullemer
-
Katharina Dreiling
-
Andreas Graur
-

Erik Kaiser
-
Haidar Kasem
-

Michael Wegert
-

Constantin Zehender
Selected Publications
Mapping Active Gene-Regulatory Regions in Human Repopulating Long-Term HSCs
Wünsche P*, Eckert ESP*, Holland-Letz T, Paruzynski A, Hotz-Wagenblatt A, Fronza R, Rath T, Gil-Farina I, Schmidt M, von Kalle C, Klein C, Ball CR, Herbst F§, Glimm H§
NRG1 Fusions in KRAS Wild-Type Pancreatic Cancer
Heining C*, Horak P*, Uhrig S*, Codo PL, Klink B, Hutter B, Fröhlich M, Bonekamp D, Richter D, Steiger K, Penzel R, Endris V, Ehrenberg KR, Frank S, Kleinheinz K, Toprak UH, Schlesner M, Mandal R, Schulz L, Lambertz H, Fetscher S, Bitzer M, Malek NP, Horger M, Giese NA, Strobel O, Hackert T, Springfeld C, Feuerbach L, Bergmann F, Schröck E, von Kalle C, Weichert W, Scholl C, Ball CR, Stenzinger A, Brors B, Fröhling S§, Glimm H§
Genetic subclone architecture of tumor clone-initiating cells in colorectal cancer
Giessler KM*, Kleinheinz K*, Huebschmann D, Balasubramanian GP, Dubash TD, Dieter SM, Siegl C, Herbst F, Weber S, Hoffmann CM, Fronza R, Buchhalter I, Paramasivam N, Eils R, Schmidt M, von Kalle C, Schneider M, Ulrich A, Scholl C, Fröhling S, Weichert W, Brors B, Schlesner M, Ball CR, Glimm H
Pan-cancer analysis of somatic copy-number alterations implicates IRS4 and IGF2 in enhancer hijacking
Weischenfeldt J*, Dubash T*, Drainas AP*, Mardin BR, Chen Y, Stütz AM, Waszak SM, Bosco G, Halvorsen AR, Raeder B, Efthymiopoulos T, Erkek S, Siegl C, Brenner H, Brustugun OT, Dieter SM, Northcott PA, Petersen I, Pfister SM, Schneider M, Solberg SK, Thunissen E, Weichert W, Zichner T, Thomas R, Peifer M, Helland A, Ball CR, Jechlinger M, Sotillo R, Glimm H§, Korbel JO§
Get in touch with us

