Division of Structural Biology of Infection and Immunity
Dr. Erec Stebbins
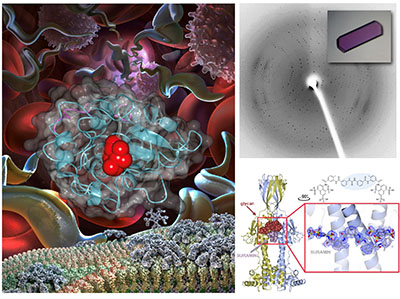
Left: Images of the African trypanosome replicating in the blood, its surface coat shown at the bottom as it switches from one VSG to the other. Right top: protein crystal and diffraction pattern. Right bottom: Structure of VSGsur bound to the anti-trypanosome drug suramin.
© dkfz.de
Immune escape variants typify long-term pathogenesis in both infectious disease and malignancy. The immune system recognizes foreign and dangerous "self" antigens, clearing cells harbouring them such as microbes and tumours. Pathogens and persistent cancers often counter immunity by varying surface antigens. We are using a controlled, bio-safe, genetically tractable model system (the African Trypanosome, T. brucei) to study long-term, immune-evasive processes in a collaborative, strong immunology program at the German Cancer Research Center. Our initial results have been paradigm shifting.
T. brucei causes sleeping sickness in humans and a related disease in animals. Central to its immune-evasion strategy is the Variant Surface Glycoprotein (VSG) that forms a unique and extremely dense coat of ~10 million molecules carpeting the surface of the organism. The VSG coat elicits a robust antibody response that the parasite evades by accessing a large genetic repertoire of divergent VSGs and switching to a new (antigenically distinct) variant, resulting in long-term infection with observable peaks and valleys of parasitemia. These oscillations are the outcome of repeated cycles of antibody generation, parasite killing, and VSG switching: a process known as antigenic variation.
Our program is to mechanistically characterize the highly diverse VSG proteins and their complexes with antibodies. Initial successes of this bottom-up approach using X-ray crystallography have dramatically altered our view of T. brucei immune evasion with discoveries such as (1) major and unexpected structural divergence in the VSGs, (2) the identification of unanticipated and immune-modulatory post-translational modifications to the VSGs, vastly expanding the epitope space, (3) antibody-VSG co-crystal structures that have for the first time mapped the epitope surface accessibility of the coat, (4) VSG function beyond mere antigenic variation (e.g., binding biologically relevant substrates), (5) demonstration that the individual VSGs elicit unusual antibody repertoires that are highly focused on immunodominant epitopes, and (6) the translational development of a novel and powerful vaccine platform based on this surface coat and critically informed by protein structural considerations.
Technicians, students, and postdocs work within this background on projects to characterize VSGs proteins (mapping their immunological diversity space) and antibody co-crystal structures with antigen. Techniques used in bench work include tissue culture, growth of eukaryotic microorganisms, large-scale protein purification and crystallization (both endogenous in the African trypanosome and recombinant, through bacterial expression), structural determination by X-ray crystallography and cryo-electron microscopy.

