Division of Molecular Genome Analysis
Prof. Dr. Stefan Wiemann
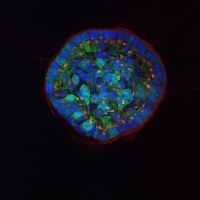
Confocal image of human breast epithelial cell line MCF10A, stably expressing ERBB2 receptor tyrosine kinase. Cells were grown in 3D (matrigel) and stained for GM130 (Golgi marker, red), phalloidin (cytoskeleton, green) and DAPI (DNA, blue). MCF10A overexpressing ERBB2 forms acini structures which are filled with viable cells. While the cells at the rim of acini are polarized, the cells in the center are not.
© dkfz.de
Cancer and many other human diseases arise from genetic aberrations that are either inherited or occur spontaneously in somatic cells. These defects cause abnormal activities of gene products and lead to malfunctioning of molecular and cellular interactions which may induce tumors and cause cancer progression. The central objective of our division is to understand the complexity of molecular mechanisms in the regulation of signaling networks and how these impact cancer development, metastasis, and drug resistance. To this end, we generate and maintain resources for large-scale experimentation, apply high-throughput functional genomics and proteomics technologies, and analyze candidate genes using in vitro as well as in vivo systems. Effects of perturbations (gene gain- and loss-of-function, miRNA, drugs) imposed on the signaling processes are experimentally tested and then computationally modeled. This generates mechanistic knowledge that is exploited to identify new diagnostic and prognostic markers as well as to develop novel strategies for therapeutic intervention. Our major focus here is on breast cancer, where we investigate protein and non-protein factors that are involved in the progression of different subtypes via their activities in interrelated signaling networks.
FUTURE OUTLOOK
We have already seen from our current data that signaling is not regulated in isolated pathways, but rather in complex networks. These networks involve different cell types within the tumor microenvironment which impact on the tumor cells and vice versa. We thus investigate the impact individual perturbations have, at different levels (DNA, RNA, protein, metabolite, phenotype
), on a variety of cellular pathways and in the context of different cell types. This should provide us with a better understanding of the connectivity between multi-layer interaction systems. Such information will be essential, for example, in identifying strategies that should help overcoming drug resistance and metastasis. While much of our current knowledge is based on in vitro experiments, we need to validate findings in vivo in order to prove their relevance. To this end, we generate and test animal models and challenge our hypotheses with data generated from patient samples. Our research is funded mostly by national (e.g., BMBF, Landesstiftung Baden-Württemberg, EKFS-Stiftung) and international (EU) agencies.

