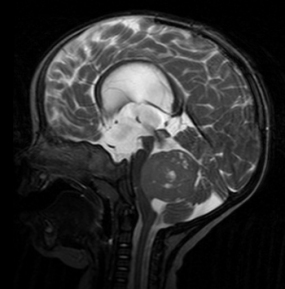
Mining the epigenome to identify likely origins of childhood brain tumor subtype
An international research team led by scientists from St. Jude Childrens Research Hospital and from German Cancer Research Center (DKFZ) mined the epigenome to discover the likely cell of origin for Group 4 medulloblastoma. The finding removes a barrier to developing more effective targeted therapies against the brain tumors most common subtype. The results appear in the scientific journal Nature.
Medulloblastoma occurs in infants, children and adults, but it is the most common malignant pediatric brain tumor. The disease includes four biologically and clinically distinct subtypes, of which Group 4 is the most common. In children, about half of medulloblastoma patients are of the Group 4 subtype. Efforts to improve patient outcomes, particularly for those with high-risk Group 4 medulloblastoma, have been hampered by the lack of accurate animal models.
Evidence from this study suggests Group 4 tumors begin in neural stem cells that are born in a region of the developing cerebellum called the upper rhomic lip. The cerebellum is the brain structure that helps coordinate movement and where medulloblastoma occurs.
Pinpointing the cell(s)-of-origin for Group 4 medulloblastoma will help us to better understand normal cerebellar development and dramatically improve our chances of developing genetically faithful preclinical mouse models. These models are desperately needed for learning more about Group 4 medulloblastoma biology and evaluating rational, molecularly targeted therapies to improve patient outcomes, said Paul Northcott, Ph.D., an assistant member of the St. Jude Department of Developmental Neurobiology. Northcott, Stefan Pfister, M.D., of the German Cancer Research Center, Heidelberg; and James Bradner, M.D., of Dana-Farber Cancer Institute, Boston, are the corresponding authors.
The discovery and other findings about the missteps fueling tumor growth came from studying the epigenome, which is the array of proteins and chemicals on or attached to DNA that work in concert to orchestrate the control of gene expression in a tissue-specific manner. DNA encodes the genome, which is the blueprint for life. The epigenome determines how these instructions in the genome are carried out in different cell types.
Researchers used the analytic tool ChiP-seq to identify and track medulloblastoma subtype differences based on the activity of epigenetic regulators. The regulators included proteins known as master regulator transcription factors. They bind to DNA sequences called enhancers and super-enhancers. The master regulator transcription factors and super-enhancers work together to regulate the expression of critical genes, such as those responsible for cell identity.
Those and other tools helped investigators identify more than 3,000 super-enhancers in 28 medulloblastoma tumors as well as evidence that the activity of super-enhancers varied by subtype. The super-enhancers switched on known cancer genes, including genes like ALK, MYC, SMO and OTX2 that are associated with medulloblastoma, researchers reported.
Knowledge of the subtype super-enhancers led to identification of the transcription factors that regulate their activity. Using computational methods, researchers applied that information to reconstruct the transcription factor networks responsible for medulloblastoma subtype diversity and identity, providing previously unknown insights into the regulatory landscape and transcriptional output of the different medulloblastoma subtypes.
The approach helped to discover and nominate Lmx1A as a master regulator transcription factor of Group 4 tumors, which led to the identification of the likely Group 4 tumor cells of origin. Lmx1A was known to play an important role in normal development of cells in the upper rhomic lip (uRL) and cerebellum. Additional studies performed in mice with and without Lmx1A in this study supported uRL cells as the likely source of Group 4 tumors.
By studying the epigenome, we also identified new pathways and molecular dependencies not apparent in previous gene expression and mutational studies, Northcott said. The findings open new therapeutic avenues, particularly for the Group 3 and 4 subtypes where patient outcomes remain inferior for the majority of affected children.
For example, researchers identified increased enhancer activity targeting the TGFb pathway. The finding adds to evidence that the pathway may drive Group 3 medulloblastoma, currently the subtype with the worst prognosis. The pathway regulates cell growth, death and other functions that are often disrupted in cancer, but its role in medulloblastoma is poorly understood.
The analysis included samples from 28 medulloblastoma tumors representing the four subtypes. Researchers believe it is the largest epigenetic study yet for any single cancer type and, importantly, the first to utilize a large cohort of primary patient tumor tissues instead of cell lines grown in the laboratory. Previous studies have suggested that cell lines may be of limited use for studying the tumor epigenome. The three Group 3 medulloblastoma cell lines used in this study reinforced the observation, highlighting significant differences in epigenetic regulators at work in medulloblastoma cell lines versus tumor samples.
Charles Lin, of Dana-Farber Cancer Institute, Boston, and Serap Erkek, of the German Cancer Research Center and the European Molecular Biology Laboratory, Heidelberg, are the first authors.
This work was principally supported by the German Cancer Aid, the German Federal Ministry of Education and Research (BMBF), the Human Frontiers Science Program; from the U.S. Department of Defense; the V Foundation and ALSAC.
Charles Y. Lin, Serap Erkek, Yiai Tong, Linlin Yin, Alexander J. Federation, Marc Zapatka, Parthiv Haldipur, Daisuke Kawauchi, Thomas Risch, Hans-Jörg Warnatz, Barbara C. Worst, Bensheng Ju, Brent A. Orr, Rhamy Zeid, Donald R. Polaski, Maia Segura-Wang, Sebastian M. Waszak, David T.W. Jones, Marcel Kool, Volker Hovestadt, Ivo Buchhalter, Laura Sieber, Pascal Johann, Lukas Chavez, Stefan Gröschel, Marina Ryzhova, Andrey Korshunov, Wenbiao Chen, Victor V. Chizhikov, Kathleen J. Millen, Vyacheslav Amstislavskiy, Hans Lehrach, Marie- Laure Yaspo, Roland Eils, Peter Lichter, Jan O. Korbel, Stefan M. Pfister, James E. Bradner und Paul A. Northcott: Active medulloblastoma enhancers reveal subgroup-specific cellular origins.
Nature 2016, DOI 10.1038/nature16546
With more than 3,000 employees, the German Cancer Research Center (Deutsches Krebsforschungszentrum, DKFZ) is Germanys largest biomedical research institute. DKFZ scientists identify cancer risk factors, investigate how cancer progresses and develop new cancer prevention strategies. They are also developing new methods to diagnose tumors more precisely and treat cancer patients more successfully. The DKFZ's Cancer Information Service (KID) provides patients, interested citizens and experts with individual answers to questions relating to cancer.
To transfer promising approaches from cancer research to the clinic and thus improve the prognosis of cancer patients, the DKFZ cooperates with excellent research institutions and university hospitals throughout Germany:
The DKFZ is 90 percent financed by the Federal Ministry of Education and Research and 10 percent by the state of Baden-Württemberg. The DKFZ is a member of the Helmholtz Association of German Research Centers.