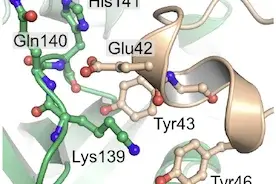
Signal Transduction in Cancer and Metabolism

Prof. Dr. Aurelio Teleman
The Teleman lab studies tissue growth and its regulation.
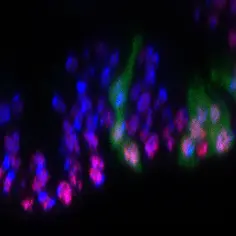
Research Synopsis
When cancer cells proliferate to form a tumor, they need to grow and to divide. Regulation of cell division (i.e. the cell cycle) has been extensively studied. In comparison, the mechanisms regulating cell growth (i.e. the accumulation of cell mass) are less well understood. The Teleman lab studies cell growth and its regulation. We study two aspects of cell growth: 1) To grow, cells need to produce biosynthetic building blocks such as nucleotides, amino acids and lipids, by activating metabolic pathways. Hence, we study cellular metabolism and how it differs in growing cells compared to non-growing cells. 2) Cells decide to grow, however, based on information coming from outside the cell, such as the presence of nutrients and growth factors. Cells receive and process this information via signaling pathways such as the insulin or TOR pathway. Therefore, we also study the signaling pathways controlling growth.
Both cellular metabolism as well as growth-signaling are highly conserved amongst animals, from the fruit fly Drosophila to humans. Powerful model systems such as Drosophila allow rapid discovery of new biology since they are simpler and more genetically tractable compared to mammals. For this reason, the Teleman lab studies cell growth using a combination of human tissue culture and Drosophila.
The aim of our work is to understand the processes regulating cell growth so that this knowledge can be used in the future for cancer therapy. Unlike other cells in the body, cancer cells have mutations that promote cell growth. Blocking this process will hopefully be a powerful method to specifically and efficiently inhibit tumor progression.
Our Team
- Show profile

Prof. Dr. Aurelio Teleman
-

Carina Binder
-

Mahrukh Butt
-

Jiayi Chu
-

Egor Diakonov
-
Thomas Erichsen
-

Dr. Fabiola Helena Garcia Cortizo
-
Katina Elisabeth Gassen
-
Ari Emiliano Hernandez Olivera
-

Fabian Hurt
-
Magda Ibrahim
-

Maria Jerome
-

David Malpartida Tous
-

Dr. Sarthak Mishra
Postdoc
-

Sandra Müller
-

Marilena Neff
-

Sabrina Neuhäuser
-

Dr. Bernhard Radlwimmer
-

Katharina Schindler
Lab Admin
-

Sibylle Schleich
-

Maximilian Schwab
-
Ilgin Selvi
-

Mara Strampe
-

Vishnuvarthan Thangarathnam
-

Roskva Torhalsdottir
-

Robert Valla
-

Dr. Ananthakrishnan Vijayakumar Maya
-

Dr. Tobias Weber
-

Dr. Yao Zhang
Selected Publications
Willnow P and Teleman AA
Figlia G, Müller S, Hagenston AM, Kleber S, Roiuk M, Quast JP, Ten Bosch N, Carvajal Ibañez D, Mauceri D, Martin-Villalba A, and Teleman AA.
Bohlen J, Fenzl K, Kramer G, Bukau B and Teleman AA.
Ahmed SMH, Maldera JA, Krunic D, Paiva-Silva GO, Pénalva C, Teleman AA* (co-corresponding) and Edgar BA*.
Schleich S, Strassburger K, Janiesch PC, Koledachkina T, Miller KK, Haneke K, Cheng Y-C, Kuechler K, Stoecklin G, Duncan KE and Teleman AA.
Demetriades C, Doumpas N, and Teleman AA
Contact
Contact Aurelio Teleman at a.teleman at dkfz.de
Get in touch with us