Functional
Genome Analysis (B070)
Deutsches
Krebsforschungszentrum,
Im Neuenheimer Feld 580
D-69120
Heidelberg,
Germany. |
 |
.
.
..
..
Systems and
Synthetic Biology have become key
research areas in the quest for understanding
living
cells. What is missing is an experimental system in which the
knowledge gained from molecular analyses and modelling could be
reproduced in an artificial
setting that is void of the risk of contamination from natural
sources. We work at setting
up a self-replicating molecular system that forms the basis toward the
establishment of an artificial biology that is entirely independent
from Nature but
identical in terms of biophysical and biochemical parameters. Utilizing
enantiomeric L-nucleotides and D-amino
acids rather than natural components, we use chemical synthesis to produce L-form nucleic acids and D-form proteins. We take advantage of these molecules in
order to
obtain basic molecular activities in a form that is a mirror-image to Nature. All parts - including co-factors such as ATP - have to
be enantiomeric to their natural counterparts and
need to be produced synthetically.
The objective is to
set up a self-replicating system, in particular aiming at enzymatically
catalysed D-protein
production. Various enzymes
and other factors are required to achieve this goal. While still some way off, such a procedure could offer a
major advance in
medicine.
Molecules produced this way could overcome several of the limitations that are currently still
inherent to
therapeutic proteins, such as
protecting them from
degradation or reducing their immunogenicity. Also, once
protein production would not be based on chemical synthesis anymore but
driven by enzymatic processes, many pieces of a mirror-image
molecular puzzle could be produced with relative ease. Thereby,
more complex
artificial
biological systems could be pieced together that - in the very long run
- may lead to the assembly of an archetypical
model of a cell.
..
In addition,
we work at the identification of factors that affect protein stability,
whether in natural or enantiomeric conformation. An emphasis is the
improvement of binder molecules, such as nanobodies. We pursue the
production of molecules,
which
exhibit superior performance, by modulating parameters such as
aggregation. This is complemented by modelling so as to
define and predict structural features that are critical for molecules'
activity.
For all projects, there are immediate practical utilities next to the
mere improvement of the basic knowledge about molecular features.
|
...
..
..
.
..
..
Establishment
of a fully synthetic mirror-image biological system (MirrorBio)


The basic motivation behind this collaborative project
was the ambition
to understand
biological processes to an extent that will permit their re-creation.
This
ability would be documented best, if functioning molecular systems
could be
established in a mirror-image, enantiomeric version, since this would
be entirely independent of any natural compounds. Instead, the
molecular components are generated by initially only chemical
processes for the production of synthetic biomolecules and their
assembly
into artificial molecular systems. Also, artificial experimental
systems will
complement current Systems Biology,
evaluating biological models experimentally.
The project was performed as an EraSynBio consortium with
partners from
three other institutions:
- Philip E.
Dawson, The Scripps Research Institute, USA
- Andreas
Plückthun, University of Zürich, Switzerland
- Jussi
Taipale, University of Helsinki, Finland
|


 |
Weidmann et al. (2016) Org. Lett. 18, 164-167. 
Olea et al. (2015) Chem. Biol. 22, 1437-1441. 
Hauser et al. (2006) Nucleic Acids Res. 34, 5101-5111. 
|
|
..
..
Exploiting
sequence and stability information for directing nanobody stability
engineering
Variable domains of camelid
heavy-chain antibodies, commonly named nanobodies, have high
biotechnological
potential. In view of their broad range of applications in research,
diagnostics and therapy, engineering their stability is of particular
interest.
Towards these ends, we analyzed the sequences and thermostabilities of
purified nanobody binders. From this data, potentially stabilizing
amino acid
variations were identified and studied experimentally. Some improved
the
stability of nanobodies by up to 6.1°C, with an average of
2.3°C. The stabilizing mechanism involves an improvement
of both
conformational stability and aggregation behavior, explaining the
different
effect on individual molecules. Other potentially stabilizing
variations
actually led to thermal destabilization of the proteins. The reasons
for this
contradiction between prediction and experiment were investigated. The
results
illustrate the potential and limitations of engineering nanobody
thermostability from a medium-throughput data set and indicate
a species-specificity of nanobody architecture. Translation of the
results is pursued in collaboration with industry partners.
..
Kunz et al. (2019) Protein Eng. Des. Sel. 32,
gzz017. 
Kunz et
al. (2018) Sci. Rep. 8,
7934.  .. ..
Kunz et al. (2015) BBA-Gen. Subjects 1861, 2196-2205. 
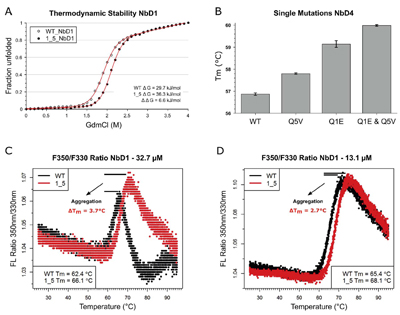
Figure legend: Mechanism
of
nanobody stabilisation by N-terminal
mutations (Q1E and Q5V). (A) The thermodynamic stability of nanobody
NbD1 and
its N-terminally mutated variant was measured in guanidinium chloride
dependent
equilibrium unfolding experiments. Unfolded protein was measured by
intrinsic
tryptophan fluorescence. Red lines represent fitted curves. (B)
Additive effect
of thermostabilisation by single and double mutations in nanobody NbD4,
measured by differential scanning fluorimetry in triplicate.
Improvements by
mutations Q5V (0.9 °C) and Q1E (2.3 °C) match the stabilisation
in the double
mutant (3.1 °C). (C, D) Tryptophan fluorescence ratio (350 nm/330
nm) for
melting nanobody NbD1 and its N-terminally mutated variant; in panel C,
a
concentration of 32.7 μM was used; in panel D, concentration was
reduced to
13.1 μM. Aggregation is indicated by a reduced amplitude of the
unfolding
transitions in the fluorescence traces and can be quantified by
comparing Tm
values of both concentration sets. Heating rate: 0.5 °C/min.
|
|
|
..
..
 |
|
Structural
Modelling:
Revised CHARMM force field parameters for iron-containing co-factors of
photosystem II
Photosystem II is a complex
protein–cofactor machinery that splits water molecules into molecular
oxygen,
protons, and electrons. All-atom molecular dynamics simulations have
the
potential to contribute to our general understanding of how photosystem
II works.
To perform reliable all-atom simulations, we need accurate force field
parameters for the cofactor molecules. We present here CHARMM bonded
and
non-bonded parameters for the iron-containing cofactors of photosystem
II that
include a six-coordinated heme moiety coordinated by two histidine
groups, and
a non-heme iron complex coordinated by bicarbonate and four histidines.
The
force field parameters presented here give water interaction energies
and
geometries in good agreement with the quantum mechanical target data.
..
Adam et al. (2018) J. Comput. Chem. 39, 7-20. 
..
Figure legend: Cover image of the Journal of Computational Chemistry,
volume 39, issue 1, on 5 January 2018. It presents artwork that is
based on the results reported in the above mentioned publication.
|
..
|