HP-F3: Epitranscriptomics, Crystallography, and Organoid Cancer Models
Type: Practical Course with Student Seminars
Part 1: Crystallography
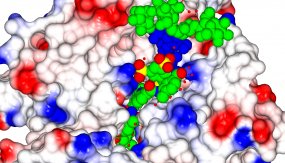
© dkfz.de
Date: 30. October 3. November 2023 (no course on 1. Nov.)
Location: DKFZ Teaching Lab and D160 Division Labs (DKFZ Main Building, 7th floor)
Hosts/Supervisors: Erec Stebbins, Johan Zeelen, Alex Hempelmann, and Monica Chandra (contacts: e.stebbins@dkfz.de, j.zeelen@dkfz.de)
Topic:
Crystallography: A tool to understand protein-protein interactions and drug discovery
Content:
- Practical introduction to crystallography
- Crystallography and cancer research
- Crystallization of a protein
- Solving the structure of a protein
Part 2: Epitranscriptomics: RNA editing and its therapeutic application
Date: 13.-17. November 2023
Location: DKFZ Teaching Lab, D150 Division Labs and Flow Cytometry Facility (DKFZ Main Building, 7th floor)
Hosts/Supervisors: Nina Papavasiliou, Riccardo Pecori, Beatrice Casati, Annette Arnold (contacts: n.papavasiliou@dkfz.de, r.pecori@dkfz.de)
Topic:
Epitranscriptomics: RNA editing and its therapeutic application
Content:
RNA editing is the most abundant epitranscriptomic modification in Metazoans. It has several essential biological functions in many organisms and humans. RNA editing is the result of the deamination of adenosine-to-inosine (A-to-I) or cytosine-to-uracil (C-to-U) leading to a base change within the RNA molecule. In this module, we explore the biological functions of RNA editing and RNA editors and how ADAR1 (one A-to-I RNA editor) can be used to manipulate genetic information for therapeutic purposes.
Functional assays to assess RNA editing activity (including tissue culture and flow cytometer approaches)
Part 3: Creating cancer models using organoids and CRISPR/Cas9 base editors
Date: 4.-8. December 2023
Location: DKFZ Teaching Lab
Host/Supervisors: Jens Puschhof, Lena Schorr, Kyanna Ouyang, Kamil Moskal, Sahana Asokan, Aurelia Saftien (contact: jens.puschhof@dkfz-heidelberg.de)
Topics:
Adult stem cells (ASCs) reside in most organs of the mammalian body, where they constantly renew tissue or replenish lost cells upon injury. This process is based on the stem cells' ability to self-renew and give rise to differentiated daughter cells. One of the most active ASCs is located at the bottom of the crypts in the intestine, from where it continuously replenishes all cell types of the epithelium every 5 days. In 2009, it was first shown that ASCs from the mouse small intestine can perform their functions in vitro as well. Upon embedding the intestinal stem cells in a gel of extracellular matrix proteins and providing them with medium containing the essential niche components, they started to self-renew, differentiate and organize into 3D structures which are referred to as organoids. These models can be propagated indefinitely, contain both ASCs and differentiated cell types and resemble the structure and function of the original organ. Organoids models are rapidly becoming a central workhorse of cancer research to study the cellular heterogeneity of tumors, test patient-specific drug efficacy, recapitulate the mutational processes active in a tumor and assess tumor-microenvironment interactions.
Content:
In this practical, organoids will be derived from mouse biopsies and engineered using CRISPR/Cas9 base editing techniques to recapitulate key mutations found in colorectal cancer. To achieve these goals, several key aspects of organoid experimentation will be introduced, including media preparation, passaging and plating of organoids, live imaging, electroporation, clonal expansion, DNA extraction and sanger sequencing to validate CRISPR success.
